High-throughput sequencing and genomes
CFAES Bioinformatics Core, Ohio State University
2026-01-29
Introduction to sequencing technologies
What do we mean by sequencing?
- Sequencing generally refers to determining the sequence of DNA, RNA, or protein fragments
- Most commonly, especially in the context of “high-throughput” sequencing,
it specifically refers to DNA sequencing
Here, we’ll only focus on DNA sequencing, keeping in mind that:
Protein sequencing involves completely different technologies
RNA can be, and usually is, sequenced as DNA as well
How is that done and why?
RNA is usually reverse transcribed to DNA (cDNA) prior to sequencing.
While it is becoming more feasible to directly sequence RNA molecules,
RNA is an unstable molecule that is easily degraded and harder to sequence.
Overview of sequencing technologies
- Sanger sequencing (since ~1985)
Sequences a single, typically PCR-amplified, DNA fragment at a time
High-throughput sequencing (HTS)
Sequences 105-109, often randomly selected, DNA fragments at a time — two types:Short-read HTS: More accurate, shorter reads (since 2005)
Long-read HTS: Less accurate, longer reads (since 2011)
These sequenced fragments of DNA are usually called reads
Sanger sequencing
- Sanger sequencing almost always starts with PCR amplification of the target DNA region —
as illustrated by Dr. Popp last week:
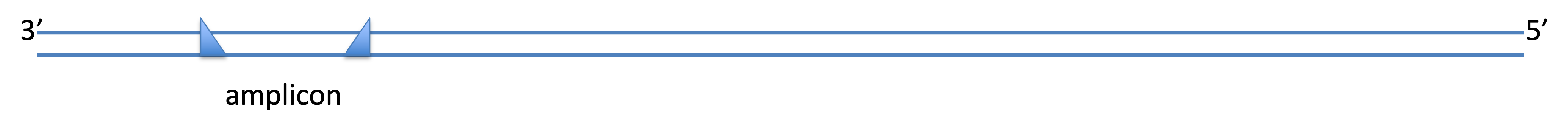
Therefore, to design primers, you must know something about the target sequence in advance — this can be highly limiting
The sequenced fragment can be up to about 800-1,000 bp
Sanger sequencing
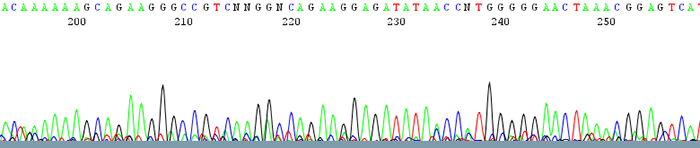
Sequencing itself is performed by synthesizing a new DNA strand with fluorescently-labeled nucleotides, using a different color for each base (
A,C,G,T)The final result is a chromatogram that can be “base-called”:
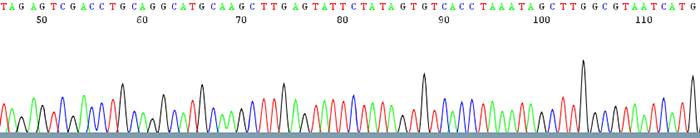
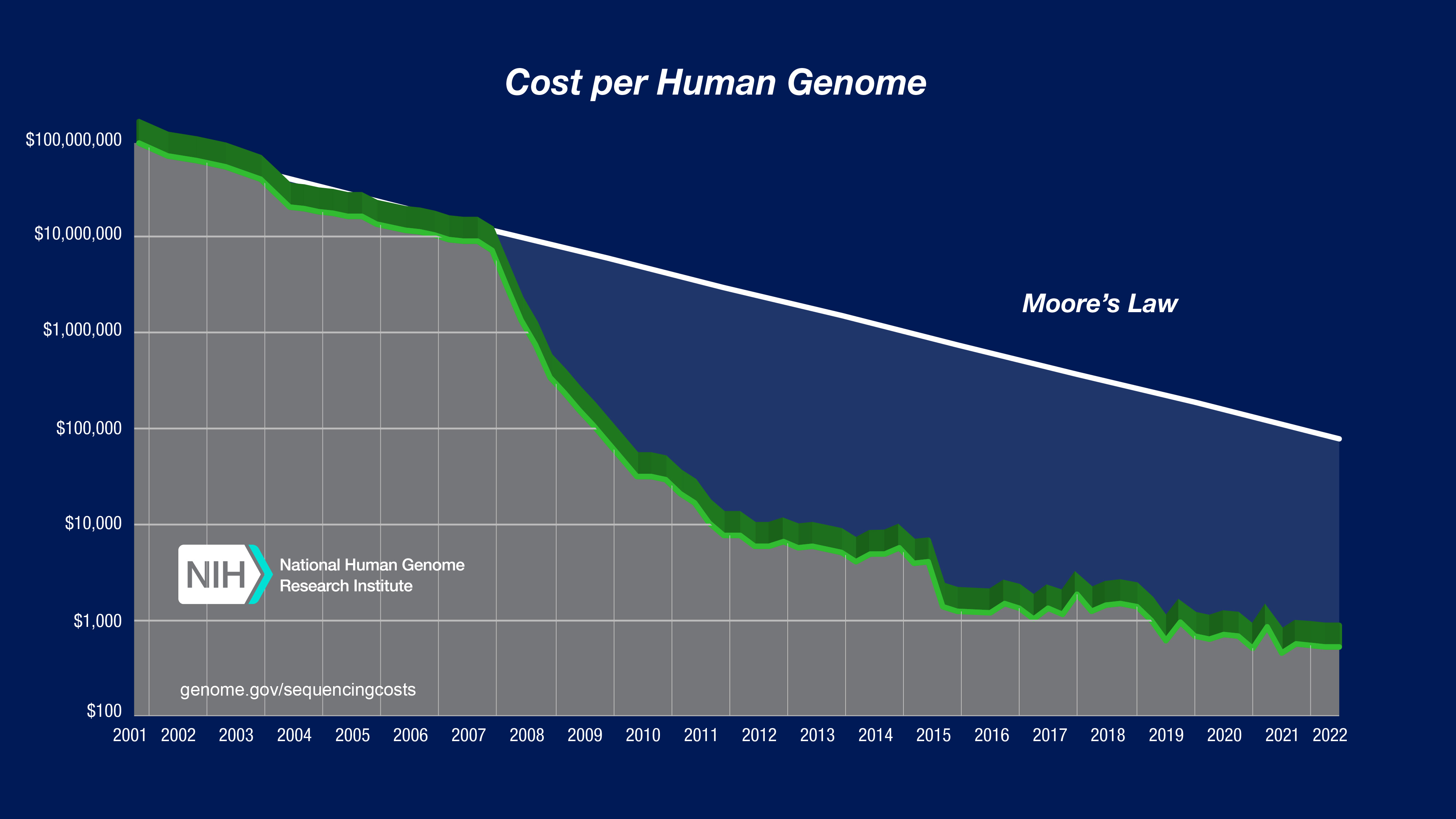
The entire human genome was sequenced with Sanger technology!
How many basepairs is that? Want to guess how much it cost to do this?
Sequencing cost through time
Present-day Sanger applications
Because HTS has much higher throughput and is much cheaper per base,
Sanger sequencing is now less widely usedBut it is not obsolete, in part because high throughput isn’t always needed
Image generated by Adobe Firefly
Present-day Sanger applications
Some present-day uses of Sanger sequencing include…
Taxonomic identification of samples
Examining genetic variation among individuals/populations
…where using one or few marker loci or candidate genes can be sufficient
High-throughput sequencing (HTS)
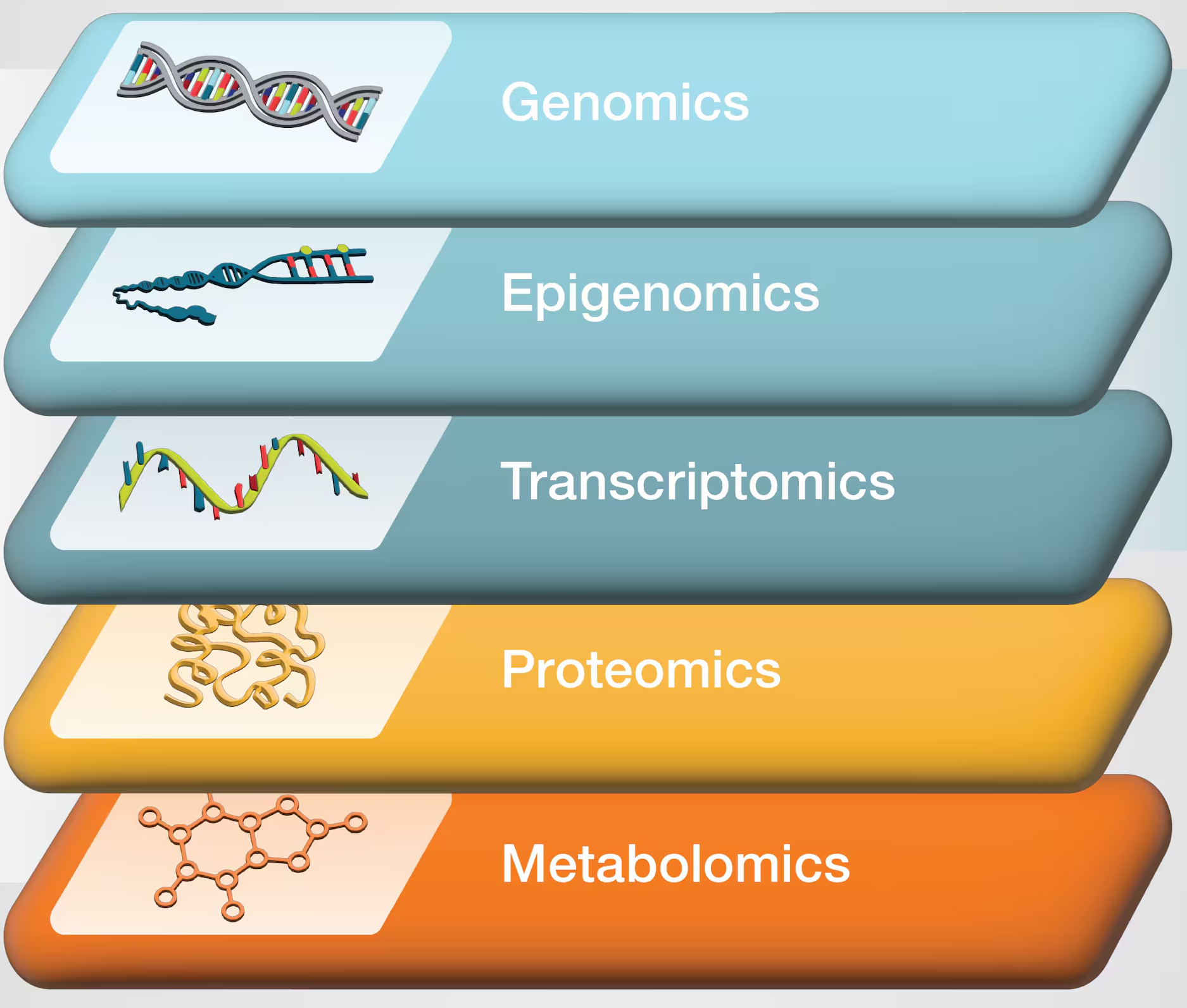
Omics
Let’s start with the big picture – HTS data underlies several of these main “omics” approaches:
Copyright ThermoFisher
The main omics data types
| Omics type | Molecule type | |
|---|---|---|
| Genomics | DNA | |
| Epigenomics | DNA modifications | High-throughput sequencing (HTS) |
| Transcriptomics | RNA | |
| Proteomics | Proteins | |
| Metabolomics | Metabolites |
What does the -omics suffix mean?
The “omics” suffix indicates the involvement of large-scale datasets —for example, “genomics” data typically spans much or all of the genome
The main omics data types
| Omics type | Molecule type | Data mainly produced by |
|---|---|---|
| Genomics | DNA | High-throughput sequencing (HTS) |
| Epigenomics | DNA modifications | High-throughput sequencing (HTS) |
| Transcriptomics | RNA | High-throughput sequencing (HTS) |
| Proteomics | Proteins | Mass Spectrometry |
| Metabolomics | Metabolites | Mass Spectrometry |
Examples of HTS applications
- Whole-genome assembly — for producing reference genomes
- Typing of SNPs and other sequence variants — for population genetics, GWAS, etc.
- Metabarcoding — for microbial community characterization
- RNA-Seq — for large-scale gene expression analysis
Does any of you work on, or is planning to work on, projects like these?
The main HTS technologies
| Short-read HTS | Long-read HTS | |
|---|---|---|
| Main companies | Illumina | Oxford Nanopore Technologies (ONT) & Pacific Biosciences (PacBio) |
The main HTS technologies
| Short-read HTS | Long-read HTS | |
|---|---|---|
| Usage | More | Less (but increasing) |
| Main companies | Illumina | Oxford Nanopore Technologies (ONT) & Pacific Biosciences (PacBio) |
| Timeline | Since 2005 — technology fairly stable | Since 2011 — still rapid development |
The main HTS technologies
| Short-read HTS | Long-read HTS | |
|---|---|---|
| Usage | More | Less (but increasing) |
| Main companies | Illumina | Oxford Nanopore Technologies (ONT) & Pacific Biosciences (PacBio) |
| Timeline | Since 2005 — technology fairly stable | Since 2011 — still rapid development |
| Read lengths | 50-300 bp | 10-100+ kbp |
| Error rates | Mostly <0.1% | 1-10% (ONT) / <0.1-10% (PacBio) |
The main HTS technologies
| Short-read HTS | Long-read HTS | |
|---|---|---|
| Usage | More | Less (but increasing) |
| Main companies | Illumina | Oxford Nanopore Technologies (ONT) & Pacific Biosciences (PacBio) |
| Timeline | Since 2005 — technology fairly stable | Since 2011 — still rapid development |
| Read lengths | 50-300 bp | 10-100+ kbp |
| Error rates | Mostly <0.1% | 1-10% (ONT) / <0.1-10% (PacBio) |
| Throughput | Higher | Lower |
| Cost per base | Lower | Higher |
Read lengths
Can you think of applications where long reads are useful?
For example:
- Genome assembly
- Read-based taxonomic identification
Can you think of applications where read length may not matter much?
For example:
- (SNP) variant analysis
- “Counting applications” such as RNA-Seq
For these, genomic locations (variant analysis) or gene identities (RNA-Seq) can be reliably inferred from as little as 25 bp
Error rates
A read’s sequence may differ from the actual DNA sequence it originated from:
- The read can have base-calling errors, missing bases, or extra bases
When you receive HTS reads, base calls have typically been made already.
Every base call is accompanied by a quality score, representing the estimated error probability.
Correcting sequencing errors
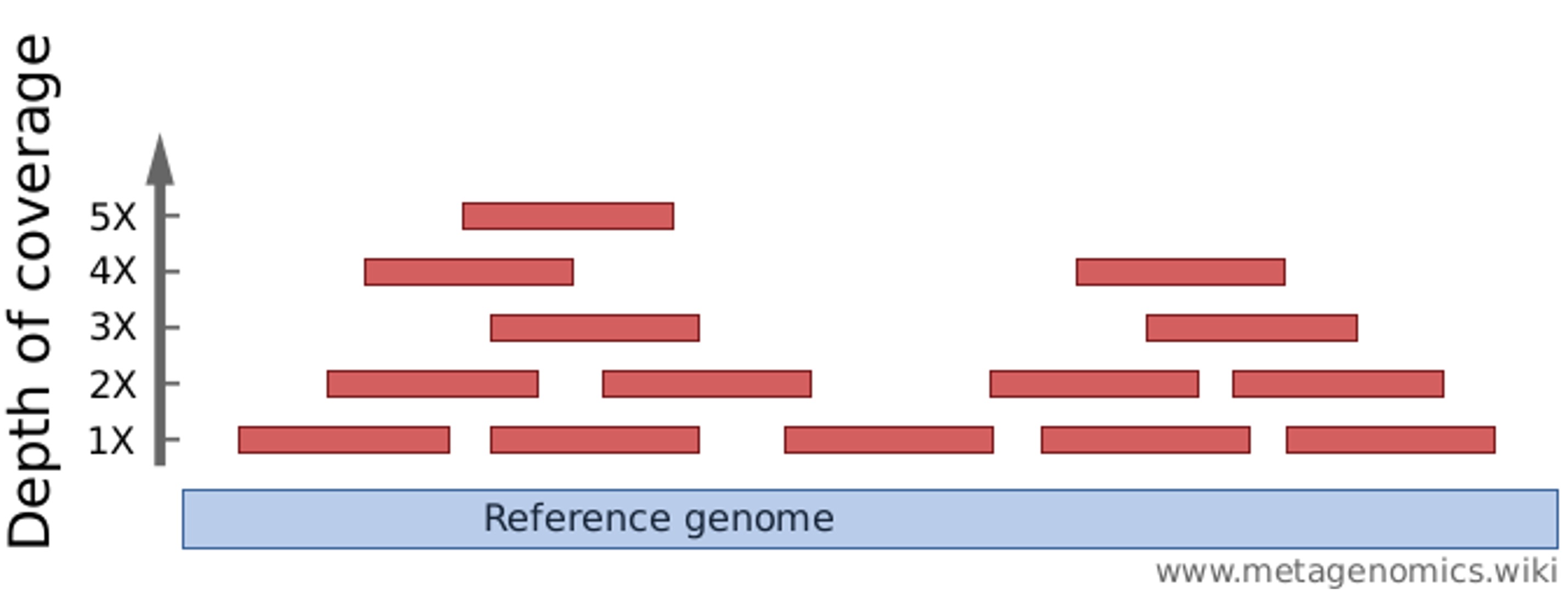
To overcome sequencing errors, every base can be sequenced multiple times –
i.e., obtaining a “depth of coverage” greater than 1:
Which natural phenomenon might complicate this effort?
Genetic variation among and (for diploid organisms) within individualsTypical depths of coverage: ~50-100x for genome assembly & 10-30x for variant typing (!)
Short-read HTS
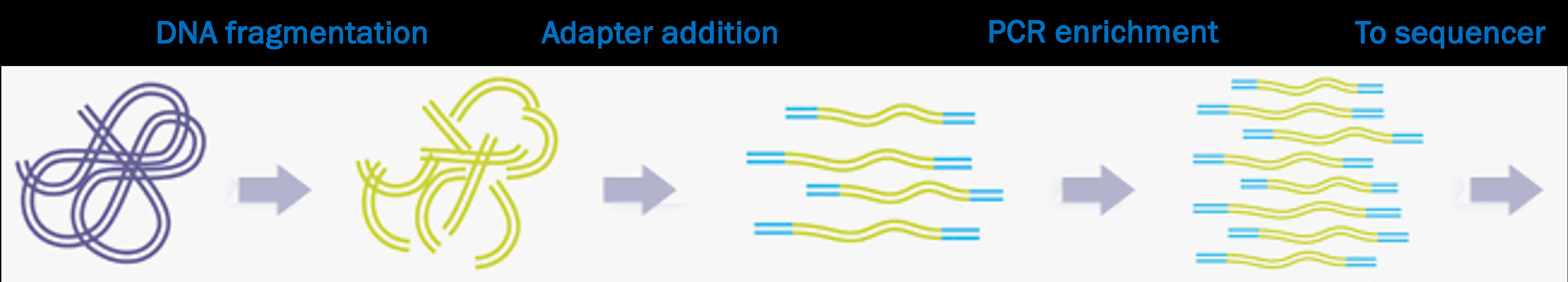
Libraries and library prep
- In a HTS context, a “library” is a collection of DNA fragments ready for sequencing
Libraries and library prep
- After library prep, each DNA fragment is flanked by several types of short sequences that together make up the “adapters”:

Multiplexing!
Adapters can include “indices” or “barcodes” to identify individual samples, so many samples can be combined (multiplexed) into a single library
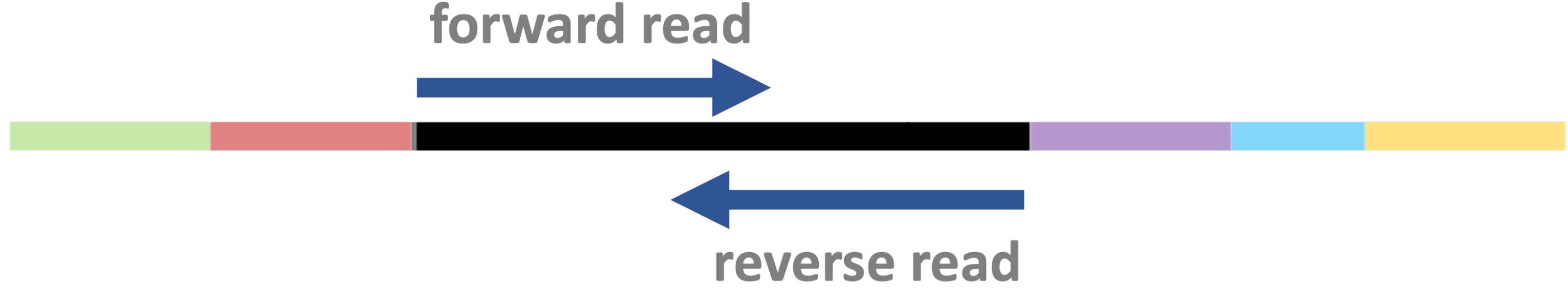
Paired-end vs. single-end sequencing
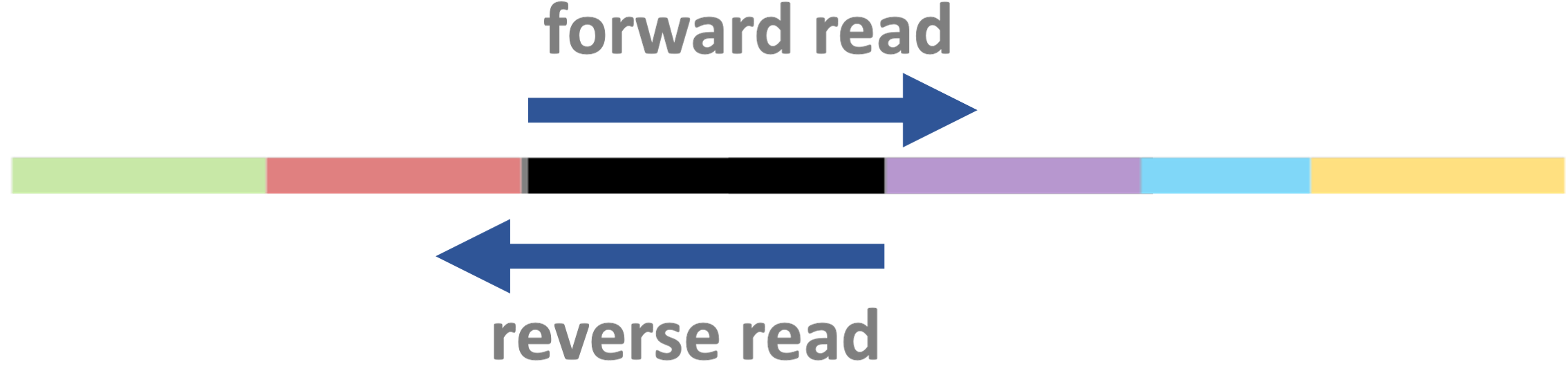
- Fragments can be sequenced from both ends as shown below —
this is called “paired-end” (PE) sequencing:
- When sequencing is instead single-end (SE), no reverse read is produced:

Fragment size variation
- DNA fragment size varies – by design and because of limited precision in size selection
What happens when a fragment is shorter than the length of a single F or R read?
“Adapter read-through”: the final bases in the resulting reads will consist of adapter sequence (these should be removed before downstream analysis)
Fragment size variation
- DNA fragment size varies – by design and because of limited precision in size selection
What happens when a fragment is shorter than the combined F + R read length?
Overlapping reads (this can be useful!):
How Illumina sequencing works
We start with fragments that have been attached to a flow cell —
this image shows just two, but millions are present simultaneously

How Illumina sequencing works
Sequencing is performed by synthesizing a new strand using fluorescently-labeled bases,
and taking a picture each time a new nucleotide is incorporated:

How Illumina sequencing works
Sequencing is performed by synthesizing a new strand using fluorescently-labeled bases,
and taking a picture each time a new nucleotide is incorporated:
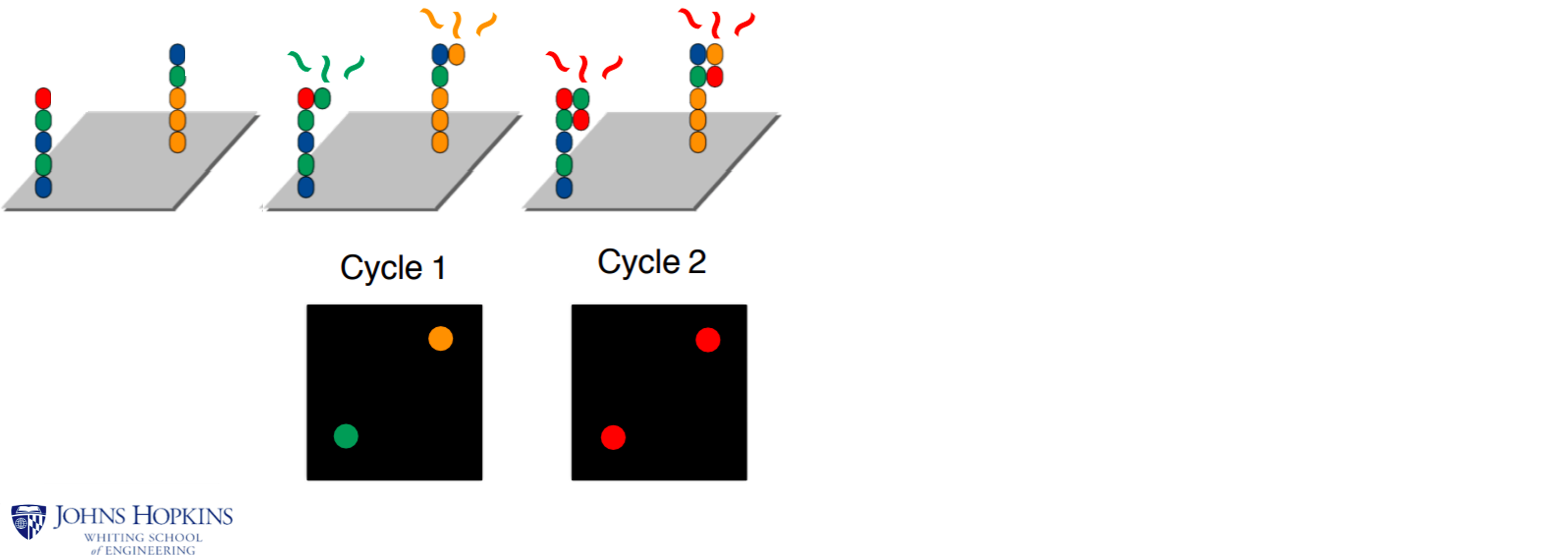
How Illumina sequencing works
Sequencing is performed by synthesizing a new strand using fluorescently-labeled bases,
and taking a picture each time a new nucleotide is incorporated:
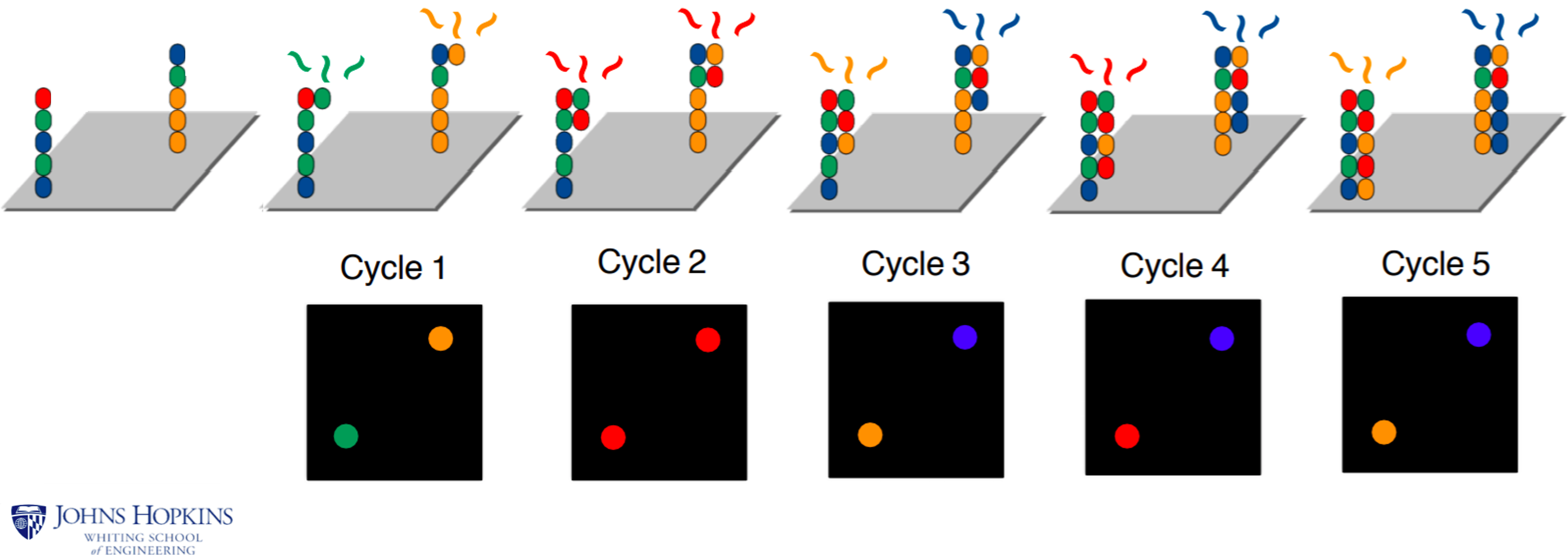
The scale of Illumina sequencing
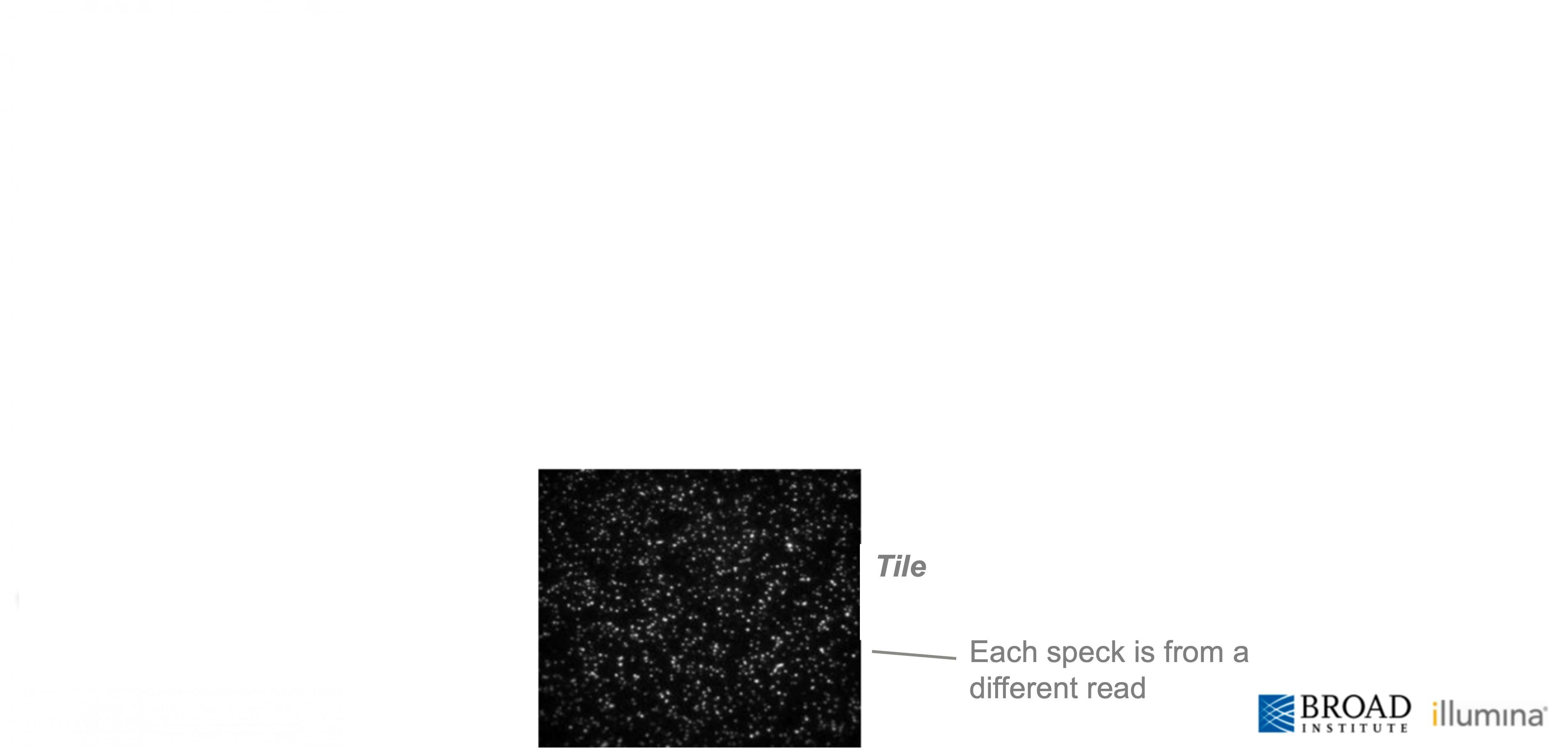

The scale of Illumina sequencing
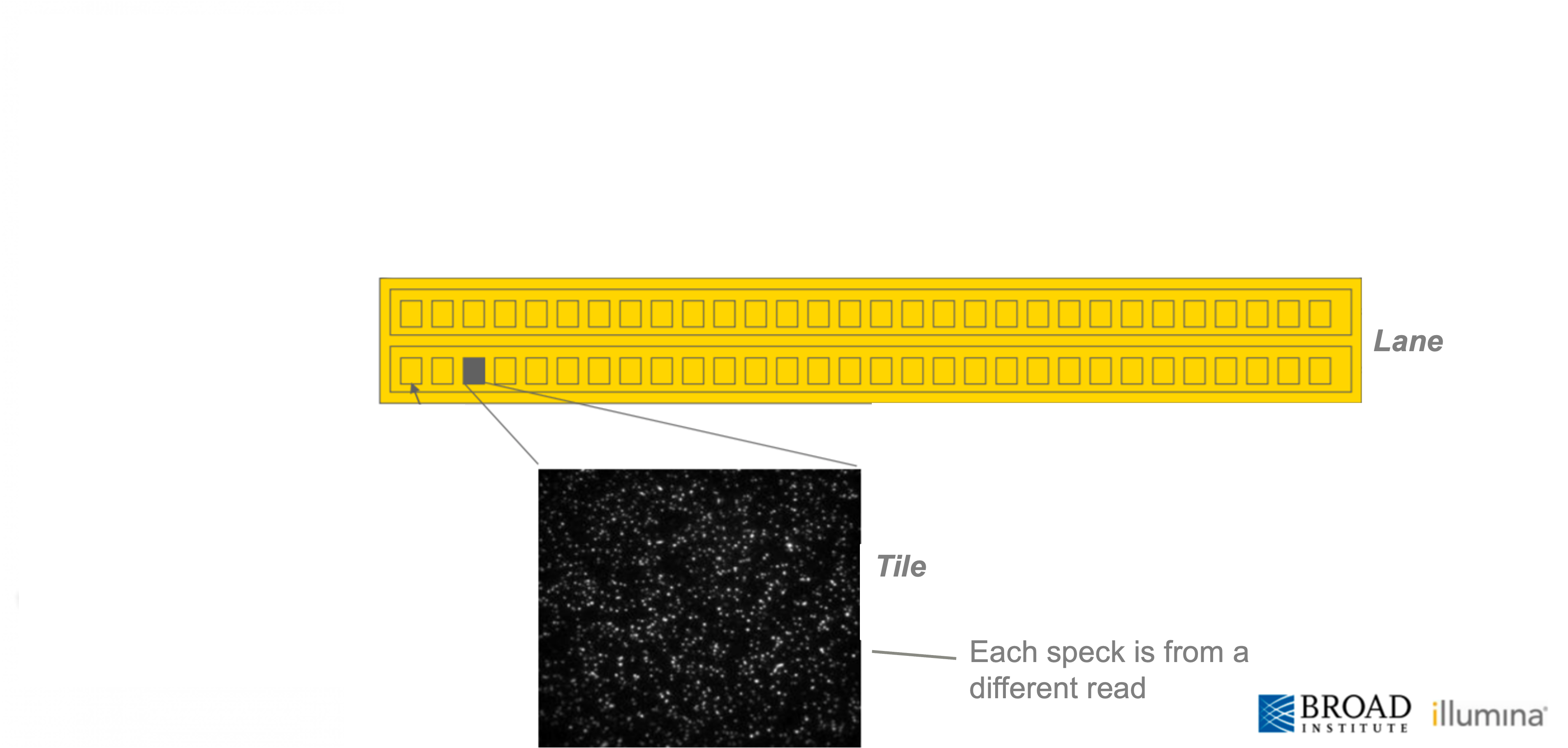
The scale of Illumina sequencing
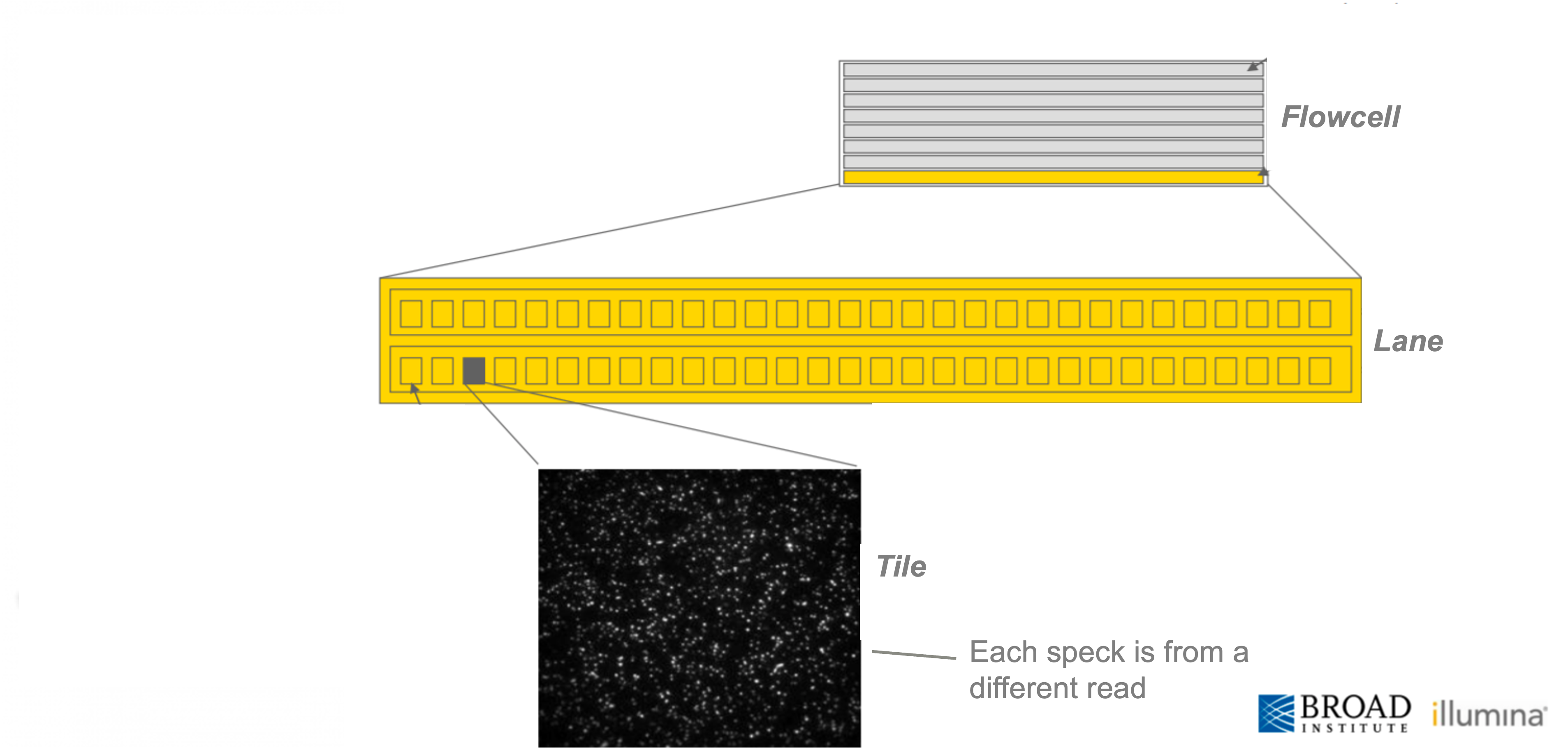
The scale of Illumina sequencing
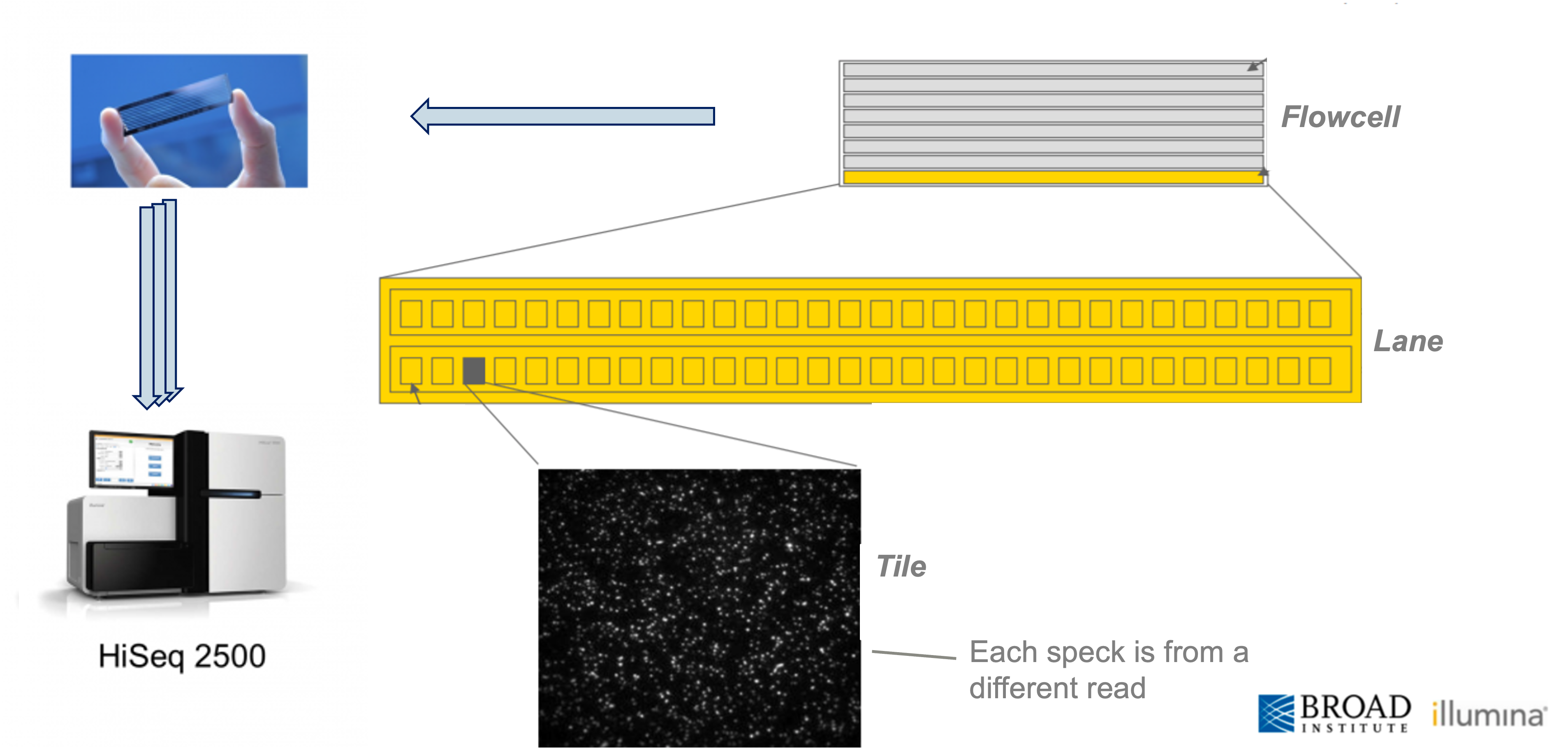
Video of Illumina technology
Reference genomes
Reference genomes
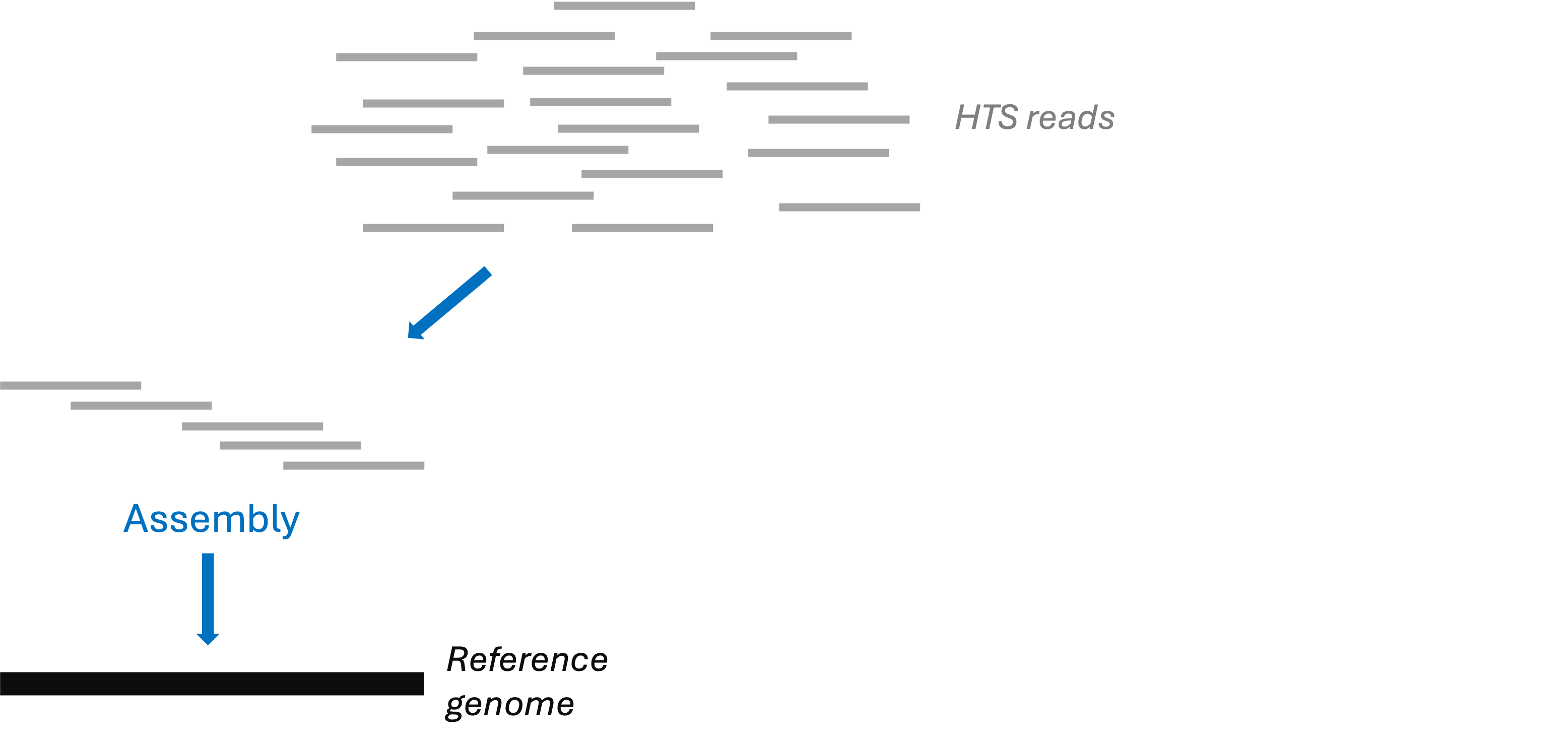
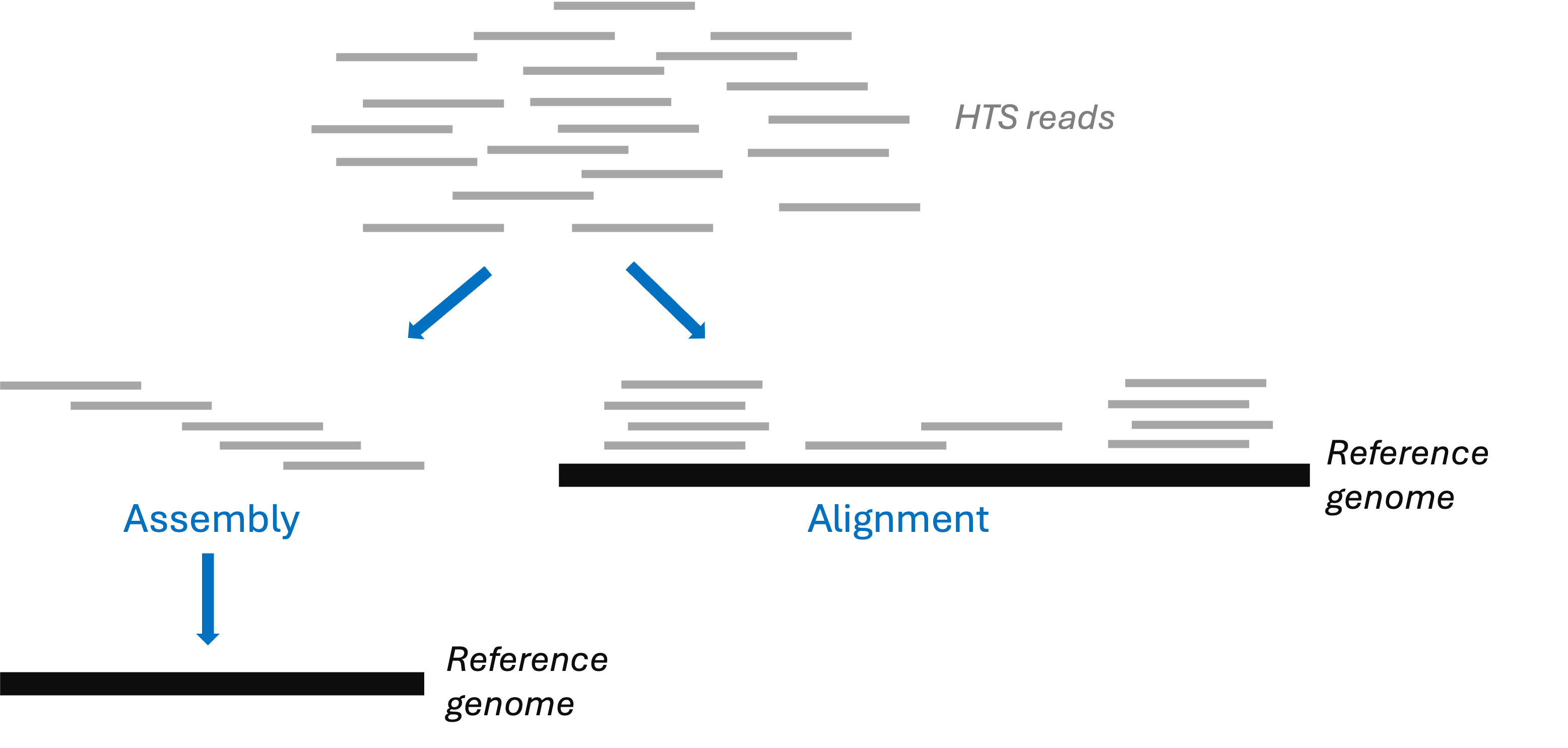
Many HTS applications either require a “reference genome” or involve its production:
Reference genomes
Many HTS applications either require a “reference genome” or involve its production:
Reference genomes
What exactly does reference genome refer to? It usually includes:
An assembly
A representation of most or all of the genome DNA sequence: the genome assemblyAn annotation
Provides e.g. locations of genes and other genomic “features” in the corresponding genome assembly, and functional information for these features
Taxonomic identity
Reference genomes are typically applicable at the species level. For example, if you work with maize, you want a Zea mays reference genome. But:
- If needed, it’s often possible to work with genomes of closely related species
- Conversely, different subspecies/lines may have their own reference genomes
There is enormous variation in genome size
And enormous growth of genomes in databases
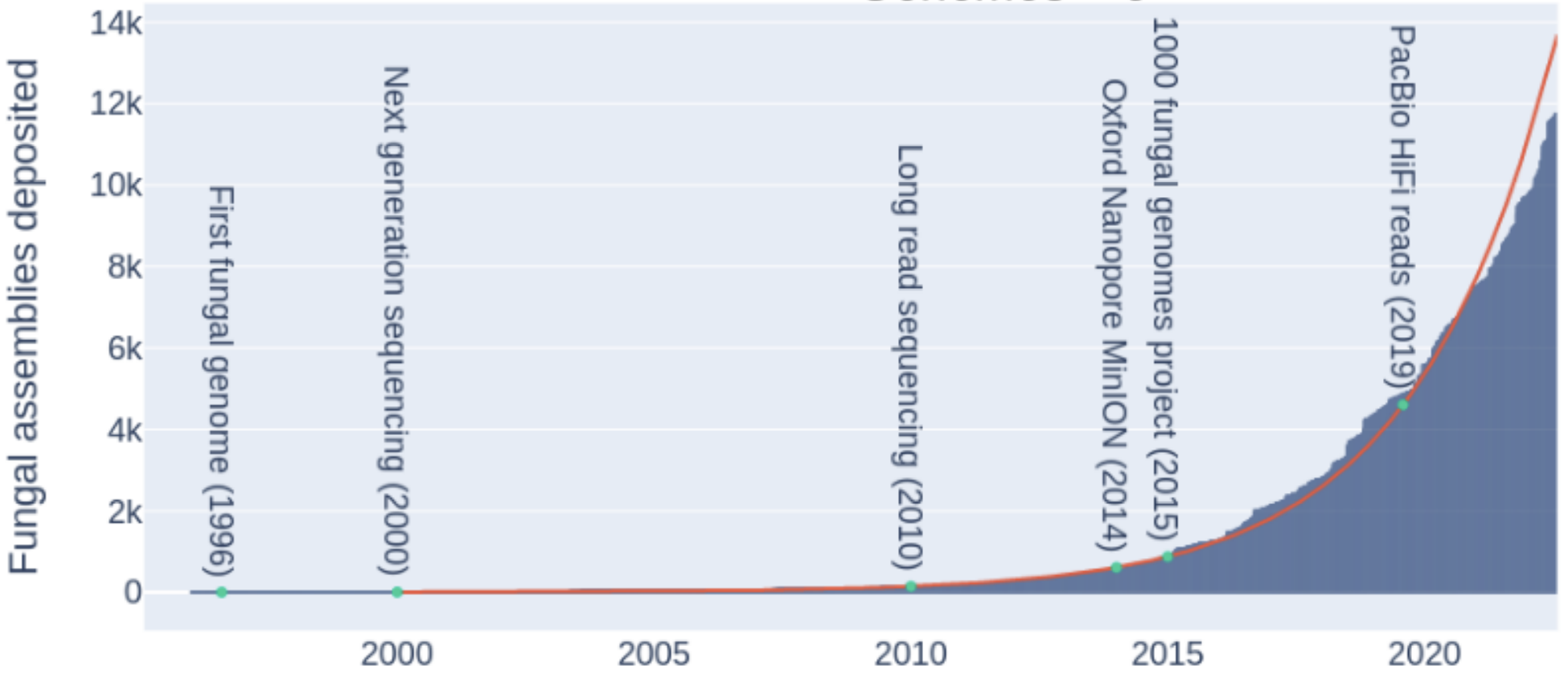
Konkel and Slot (2023)
Genome assemblies
- Besides being produced at higher rates as shown on the previous slide,
assemblies keep getting better with increasing usage & quality of long-read HTS
- Still, many assemblies instead consist of –often 1000s of– fragments (contigs and scaffolds)
How is this data stored?
Both genome assemblies and annotations are typically saved in a single text file each — we’ll explore these in tomorrow’s lab
Recap
You’ve learned…
That high-throughput sequencing (HTS) enables DNA sequencing at much larger scales than was previously possible
That HTS underlies several major “omics” fields
How short-read and long-read HTS have different strengths and weaknesses
About libraries and the technology underlying short-read sequencing
- That reference genomes are essential for many HTS applications
Looking forward
The Garrigós et al. 2025 dataset
The labs this and next week are organized around the data set from Garrigós et al. (2025):
This paper uses paired-end Illumina RNA-Seq data to study gene expression in Culex pipiens mosquitos infected with two different malaria-causing Plasmodium protozoans.
Tomorrow’s lab
- You will first learn about what kind of computing environment is commonly used for analyzing high-throughput sequencing (HTS) data
- Then, you’ll work in this environment to explore an HTS dataset, checking out HTS read and reference genome files, and performing read quality control
Next week’s content
- In the lecture, you will learn how RNA-Seq is used to study gene expression at scale
- In the lab, you will perform differential gene expression analysis on the Culex pipiens RNA-Seq dataset
Bonus: More on HTS applications
Examples of HTS applications
We talked about these HTS applications:
- Whole-genome assembly — for producing reference genomes
- Typing of SNPs and other sequence variants — for population genetics, GWAS, etc.
- Metabarcoding — for microbial community characterization
- RNA-Seq — for large-scale gene expression analysis
In each of of these applications:
- What part(s) of the genome are you sequencing?
- Who are you sequencing (how many individuals/species)?
- How are the sequences used?
Examples of HTS applications
| Appl. | What | Who | How |
|---|---|---|---|
| Whole-genome assembly | The whole genome! 😃 | A single individual | “Overlap” reads into larger fragments |
| Variant typing | |||
| 16S Metabarcoding | |||
| RNA-Seq |
Examples of HTS applications
| Appl. | What | Who | How |
|---|---|---|---|
| Whole-genome assembly | Whole genome 😃 | A single individual | “Overlap” reads into larger fragments |
| Variant typing | Whole genome or specific (e.g. exome) or random (GBS/RAD) regions |
Multiple individuals | For each site & individual, determine the variant state (allele) |
| 16S Metabarcoding | |||
| RNA-Seq |
Examples of HTS applications
| Appl. | What | Who | How |
|---|---|---|---|
| Whole-genome assembly | Whole genome 😃 | A single individual | “Overlap” reads into larger fragments |
| Variant typing | Whole genome or specific (e.g. exome) or random (GBS/RAD) regions |
Multiple individuals | For each site & individual, determine the variant state (allele) |
| 16S Metabarcoding | A specific locus (e.g. 16S) | Multi-species samples — soil, gut contents, etc. |
Assign reads a taxonomic identity and count |
| RNA-Seq |
Examples of HTS applications
| Appl. | What | Who | How |
|---|---|---|---|
| Whole-genome assembly | Whole genome 😃 | A single individual | “Overlap” reads into larger fragments |
| Variant typing | Whole genome or specific (e.g. exome) or random (GBS/RAD) regions |
Multiple individuals | For each site & individual, determine the variant state (allele) |
| 16S Metabarcoding | A specific locus (e.g. 16S) | Multi-species samples — soil, gut contents, etc. |
Assign reads a taxonomic identity and count |
| RNA-Seq | Whole transcriptome | Multiple indivuals (& multiple conditions) |
Assign reads a gene identity and count |