Transcriptomics with RNA-Seq
CFAES Bioinformatics Core, OSU
2026-02-05
Why study gene expression?
Gene expression is highly dynamic
Unlike an organism’s genome content, the expression of genes is highly dynamic – it varies:
Across time and space
Qualitatively (which genes are expressed) and especially quantitatively (how much of each gene is expressed)
Gene expression is functionally important
Considering:
That protein production tells us about the activity of biological functions,
and the molecular mechanisms underlying those functionsThat it is easier to measure transcript (mRNA) than protein abundance
The central dogma

… gene expression can be used as a proxy for protein expression to make functional inferences
Caveat: the correlation between mRNA and protein levels is rather imperfect –
see e.g. Ponomarenko et al. (2023).
Studying gene expression
For example, gene expression can be quantified to associate genes and genetic pathways with:
- Phenotypic responses e.g. to environmental conditions
- Differences between groups (treatments, genotypes, sexes, tissues, etc.)
So, studying gene expression can identify candidate genes underlying phenotypes –
these can next by functionally validated using experiments like knock-outs.
And further down the line, understanding molecular basis of phenotypes has all sorts of uses!
E.g, in an agricultural context, to manipulate phenotypes such as resistance, pathogenicity, and yield.
How to quantify gene expression?
- Techniques like qPCR/ddPCR can quantify expression of a few genes at a time
- Useful for hypothesis-driven research on specific genes
- Not suitable for discovery-based research on many genes
- With transcriptomics, expression of thousands of genes is quantified simultaneously
- It was already common practice before HTS: with microarrays
- But is now dominated by the HTS-based method RNA-Seq
Introduction to RNA-Seq
What is RNA-Seq?
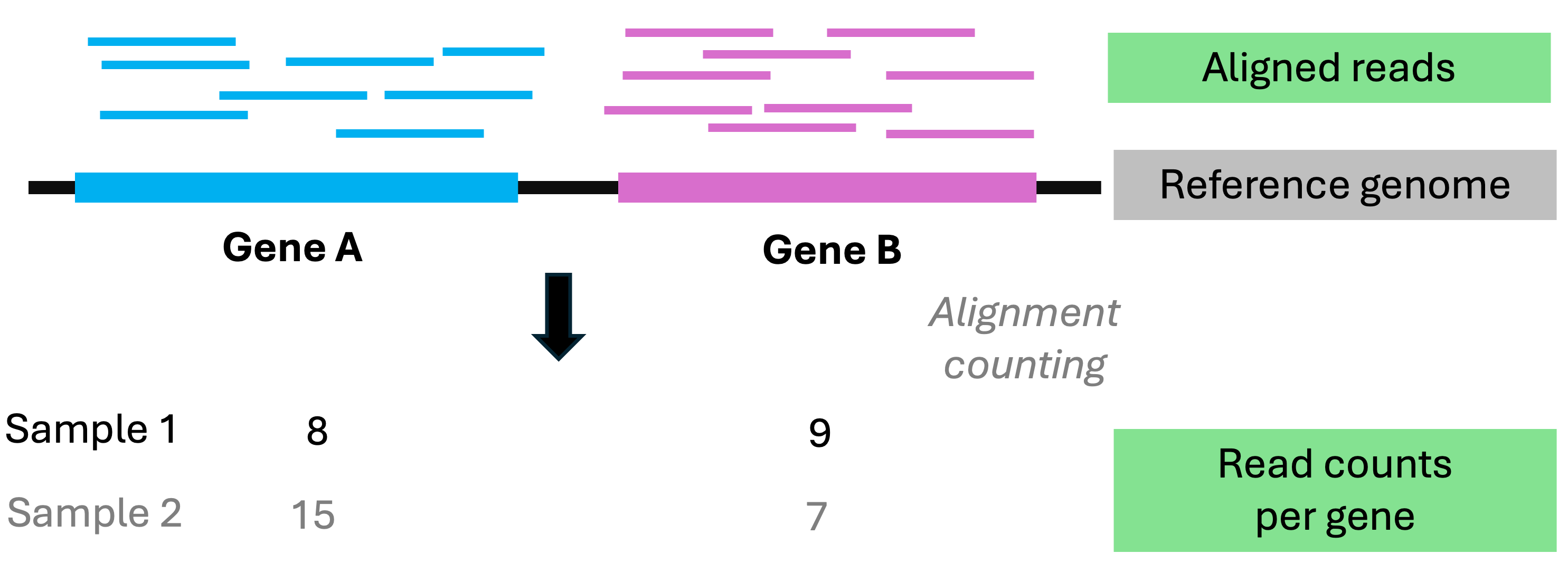
To estimate gene expression levels genome-wide, RNA-Seq takes a brute-force approach by randomly sequencing of millions of RNA fragments per sample
The resulting reads can be assigned a gene of origin,
and the core idea is that a gene’s read count reflects that gene’s expression level

How can we tell which gene each read originates from?
Most commonly by aligning the reads to a reference genome.What is RNA-Seq?
To estimate gene expression levels genome-wide, RNA-Seq takes a brute-force approach by randomly sequencing of millions of RNA fragments per sample
The resulting reads can be assigned a gene of origin,
and the core idea is that a gene’s read count reflects that gene’s expression level

RNA-Seq is a very widely used technique —
it constitutes the most common usage of high-throughput sequencing
The most common type of RNA-Seq
We’ll focus on the most common type of RNA-Seq, which:
- Does not sequence RNA directly, but first reverse transcribes RNA to cDNA
- Attempts to sequence only mRNA, avoiding non-coding RNAs (“mRNA-Seq”)
- Does not distinguish between RNA from different cell types (“bulk RNA-Seq”)
- Uses short reads (≤150 bp) that do not cover full transcripts but do uniquely ID genes
- Uses a reference genome for read alignment (“reference-based RNA-Seq”)
When I refer to RNA-Seq from now on, this specific type of RNA-Seq is implied.
Side note: Types of RNA
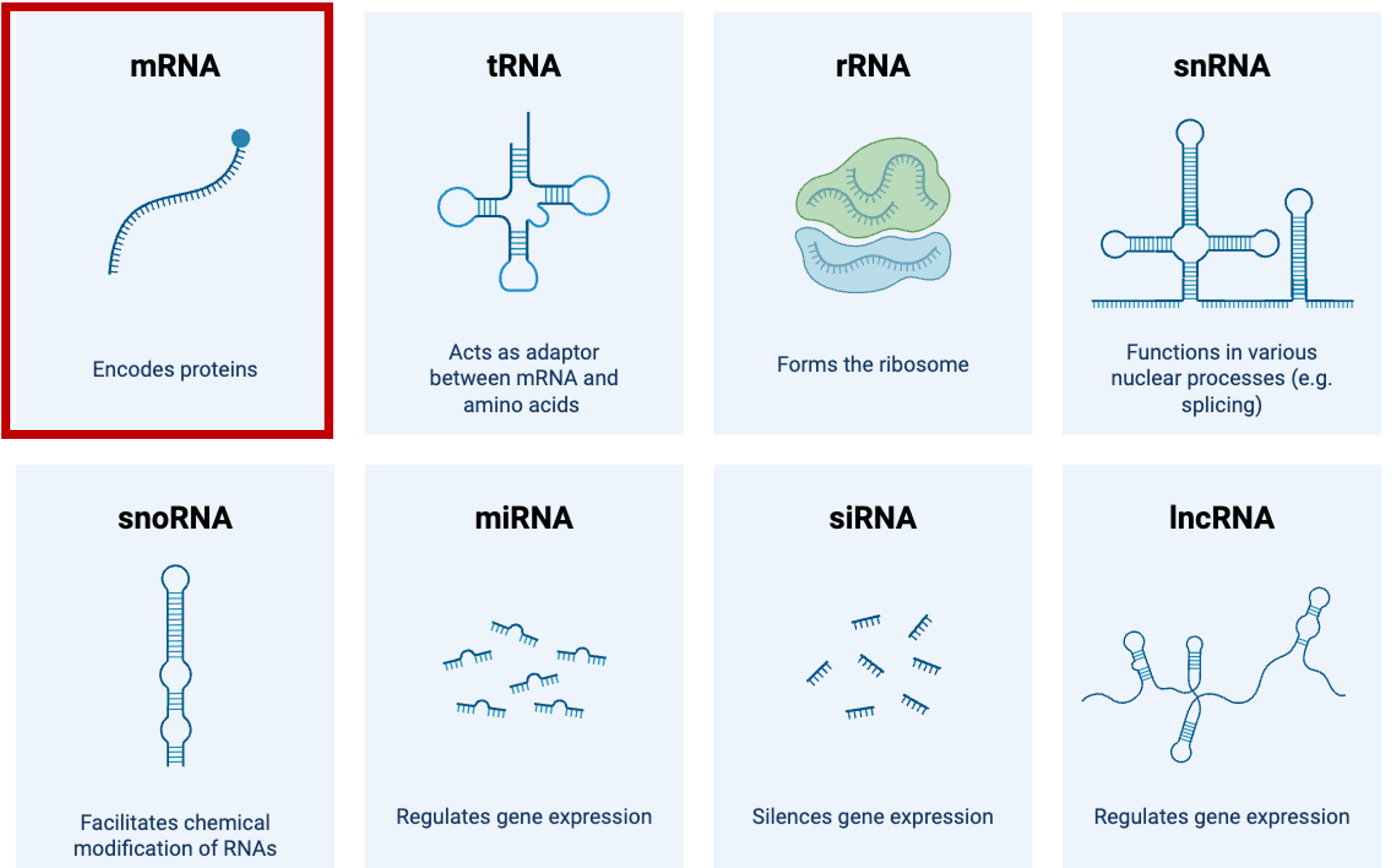
From BioRender
RNA-Seq project examples
RNA-Seq is the most common data type I help analyze in my role.
For a taste of what it’s used for in CFAES and beyond —
the following projects aimed to identify genes & pathways differing between:
Soybean cultivars in response to Phytophtora sojae inoculation (Dorrance lab, PlantPath)
Response of maize to infection with Pantoea with and without an effector gene (Mackey lab, HCS)
Mated and unmated mosquitos (Sirot lab, College of Wooster)
Tissues of the ambrosia beetle and its symbiotic fungus (Ranger lab, USDA)
Diapause-inducing conditions for two pest stink bug species (Michel lab, Entomology)
Pig coronaviruses with vs. without an experimental insertion (Wang lab, CFAH)
Human carcinoma cell lines with vs. without a manipulated gene (Cruz lab, CCC)
And to improve the annotation of a nematode genome assembly (Taylor lab, PlantPath)
Stage I: Experimental design
Experimental design: treatments/groups
RNA-Seq typically compares groups of samples defined by differences in:
Treatments — e.g. different host plants, temperature, diet, mated/unmated
Organismal variants — e.g. ages/developmental stages, sexes, subspecies
Tissues
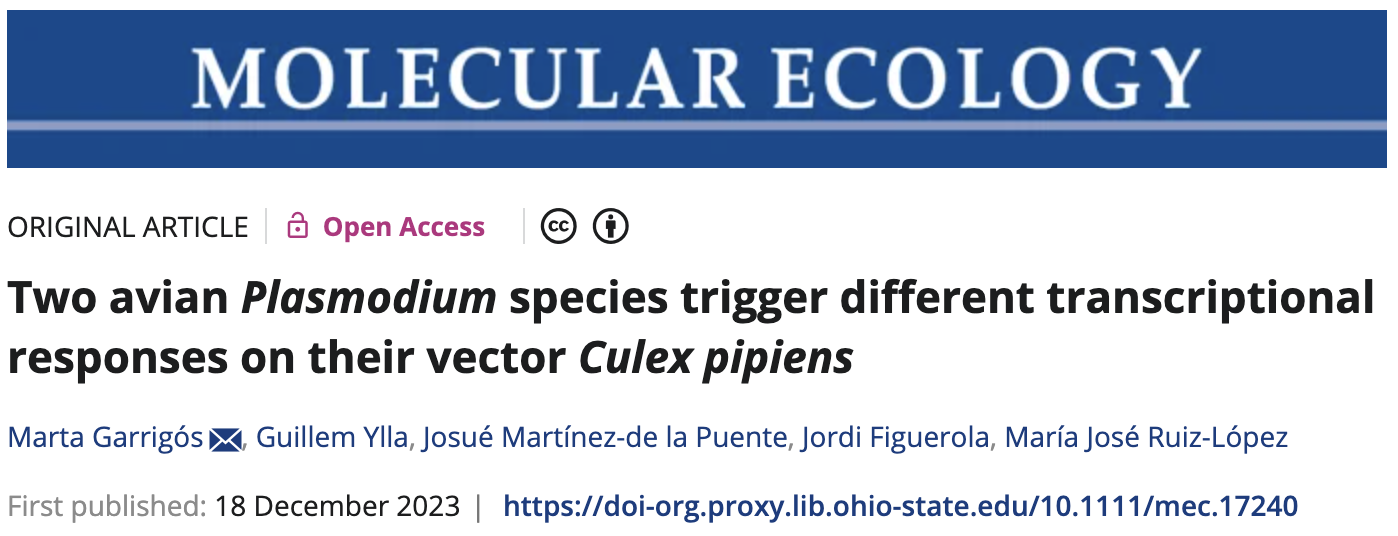
The Garrigós et al. experimental design
Garrigós et al. (2025)
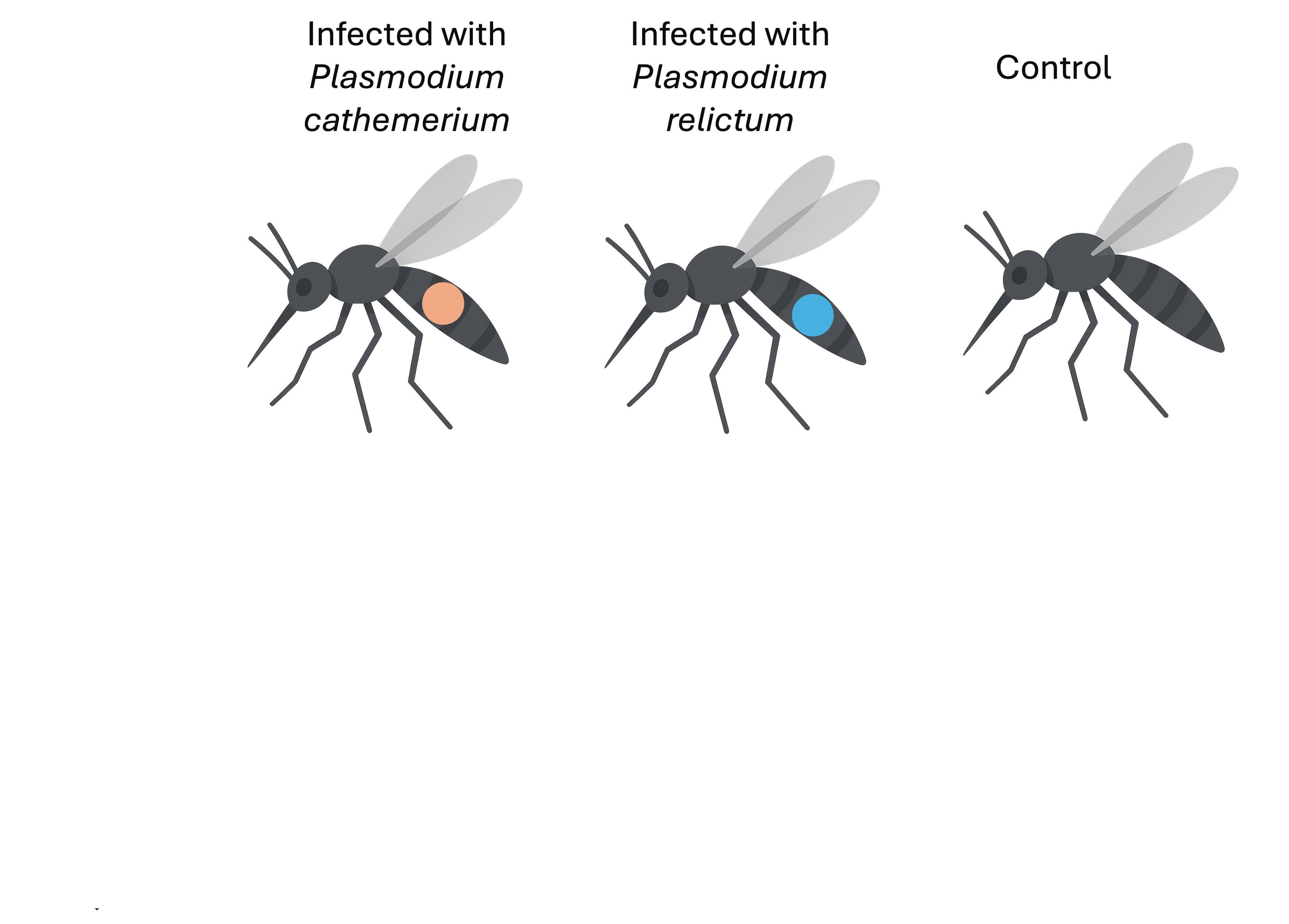
Culex pipiens mosquitos infected with malaria-causing Plasmodium protozoans:
- Plasmodium relictum – higher virulence, lower tranmission rate
- Plasmodium cathemerium – lower virulence, higher tranmission rate
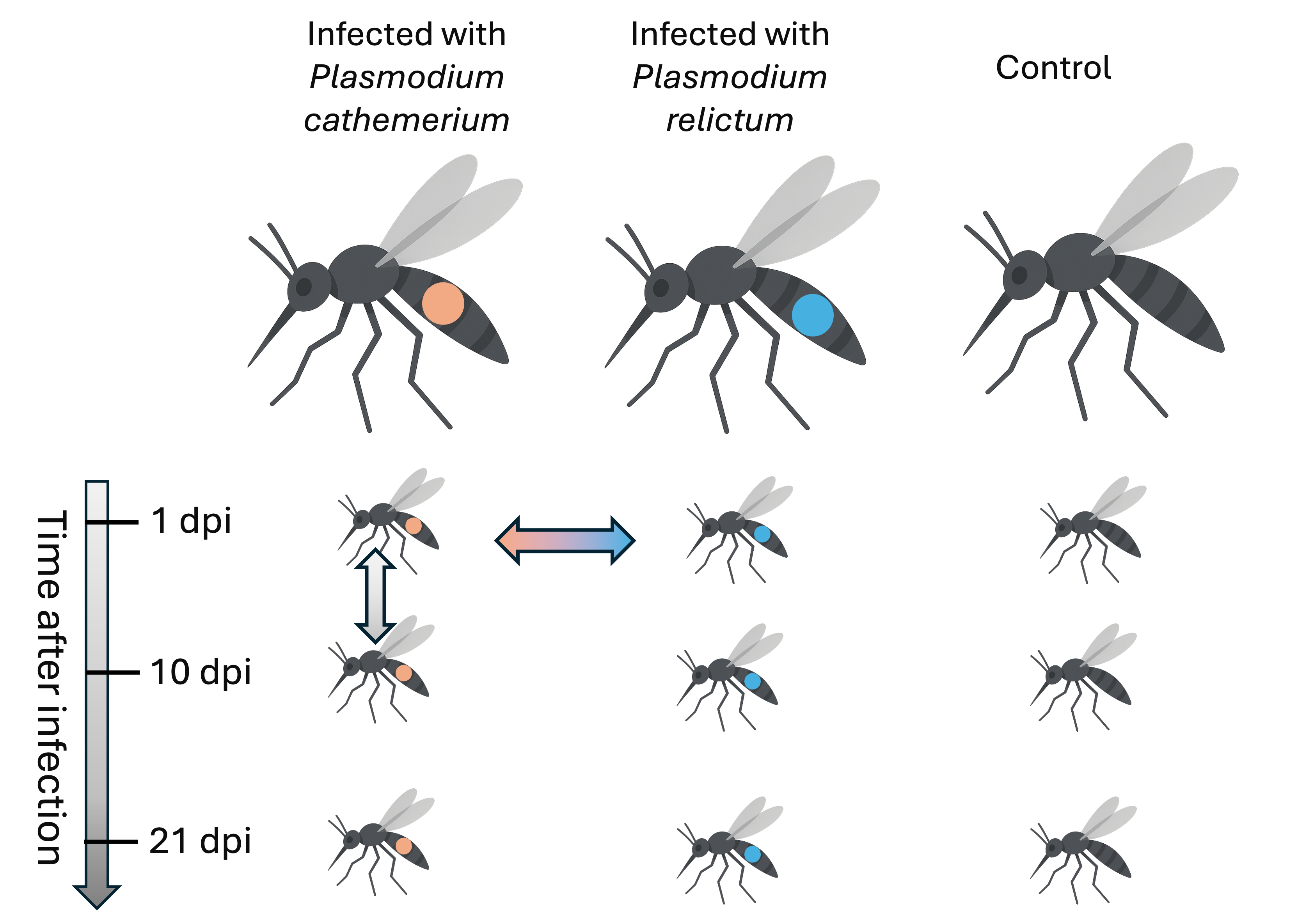
The Garrigós et al. experimental design
The Garrigós et al. experimental design
The Garrigós et al. experimental design
Experimental design
With this experimental design, would it be appropriate to sequence 9 samples total?
No – see the next slideExperimental design: Biological replicates
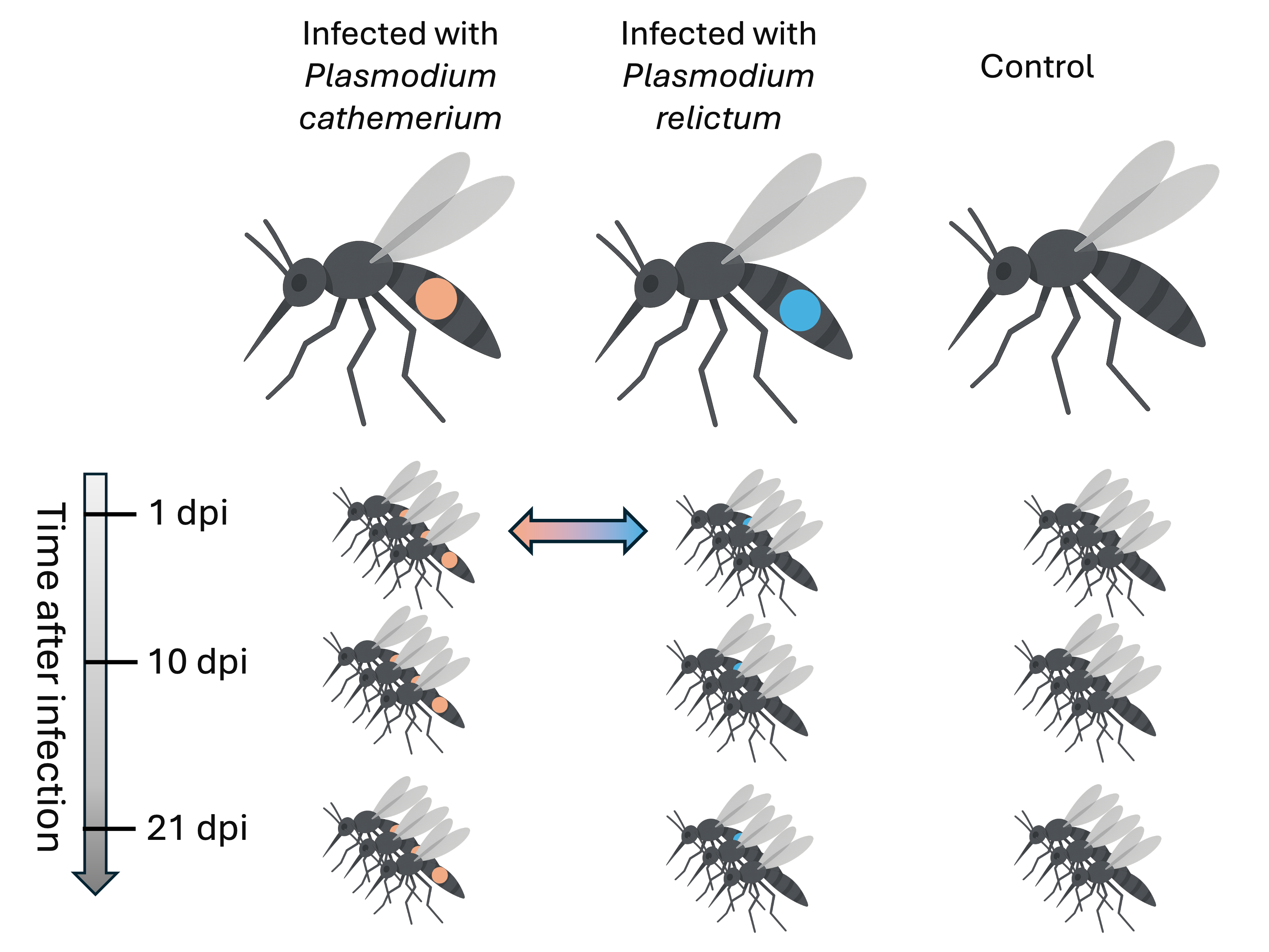
To make statistically supported conclusions about expression differences,
we need biological replication (at least 3-5 samples per group):
Stage II: From samples to reads
From samples to reads
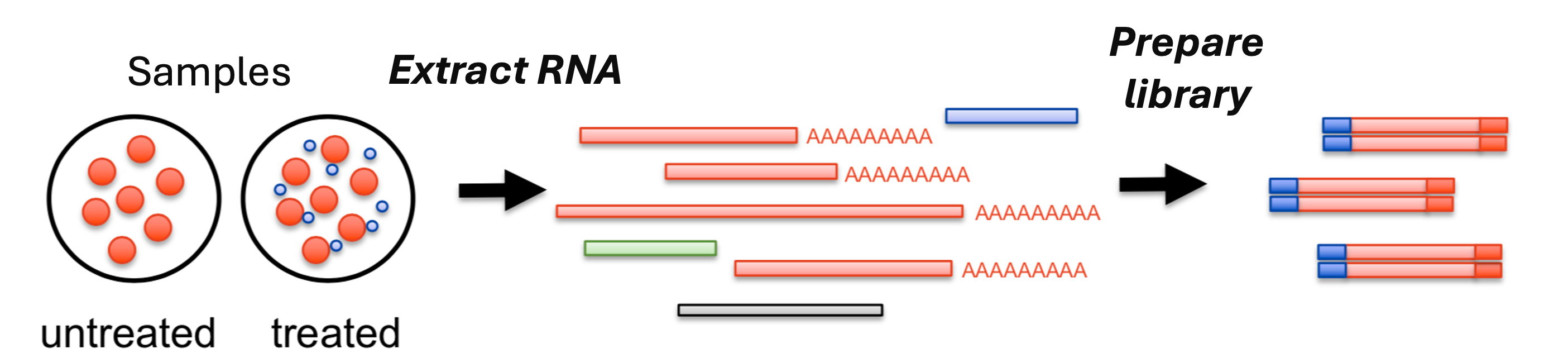
Which type of RNA do the red bars followed by AAAAAAA represent?
These are the mRNAs, which have poly-A tails (AAAAAA...)
From samples to reads

We want mRNAs but these often make up only a few percent of RNAs!
The two main ways to select for mRNAs are poly-A selection and ribo-depletion.
As mentioned in the previous lecture:
Library preparation is typically done by sequencing facilities / companies
Many samples can be “multiplexed” into a single (RNA-Seq) library
From samples to reads
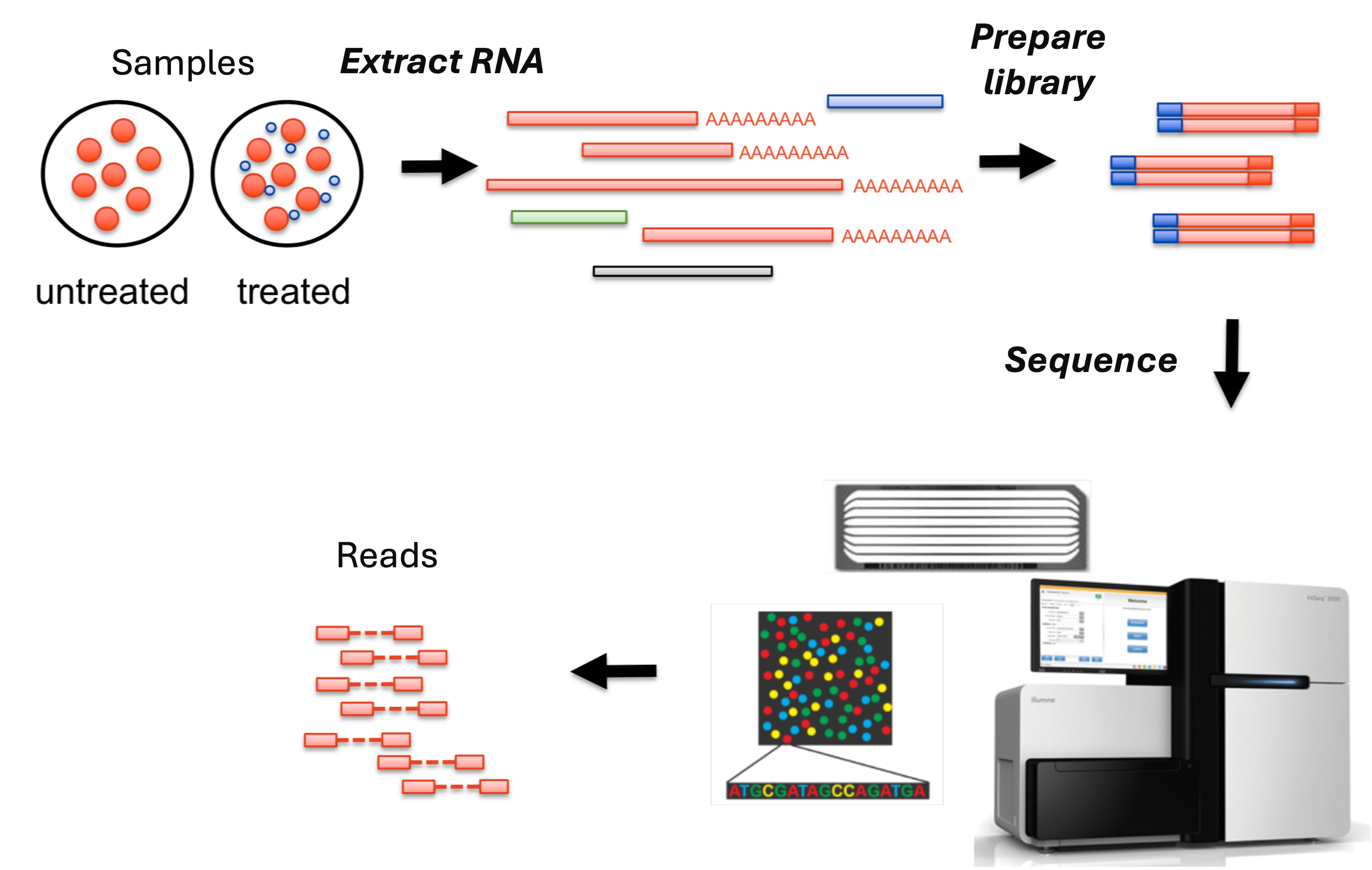
Modified after https://sydney-informatics-hub.github.io
How much sequencing is needed?
- Guidelines highly approximate — required amount depends not just on transcriptome size but also on expression level distribution, expression levels of genes of interest, etc.
- Typical recommendations are 20-50 million reads per sample in eukaryotes
Stage III: From reads to counts
Overview of RNA-Seq data analysis
The analysis of RNA-Seq data can be divided into two main parts:
- From reads to counts: from the raw reads, produce a count table
- From counts to conclusions: analyze the count table to draw biological conclusions
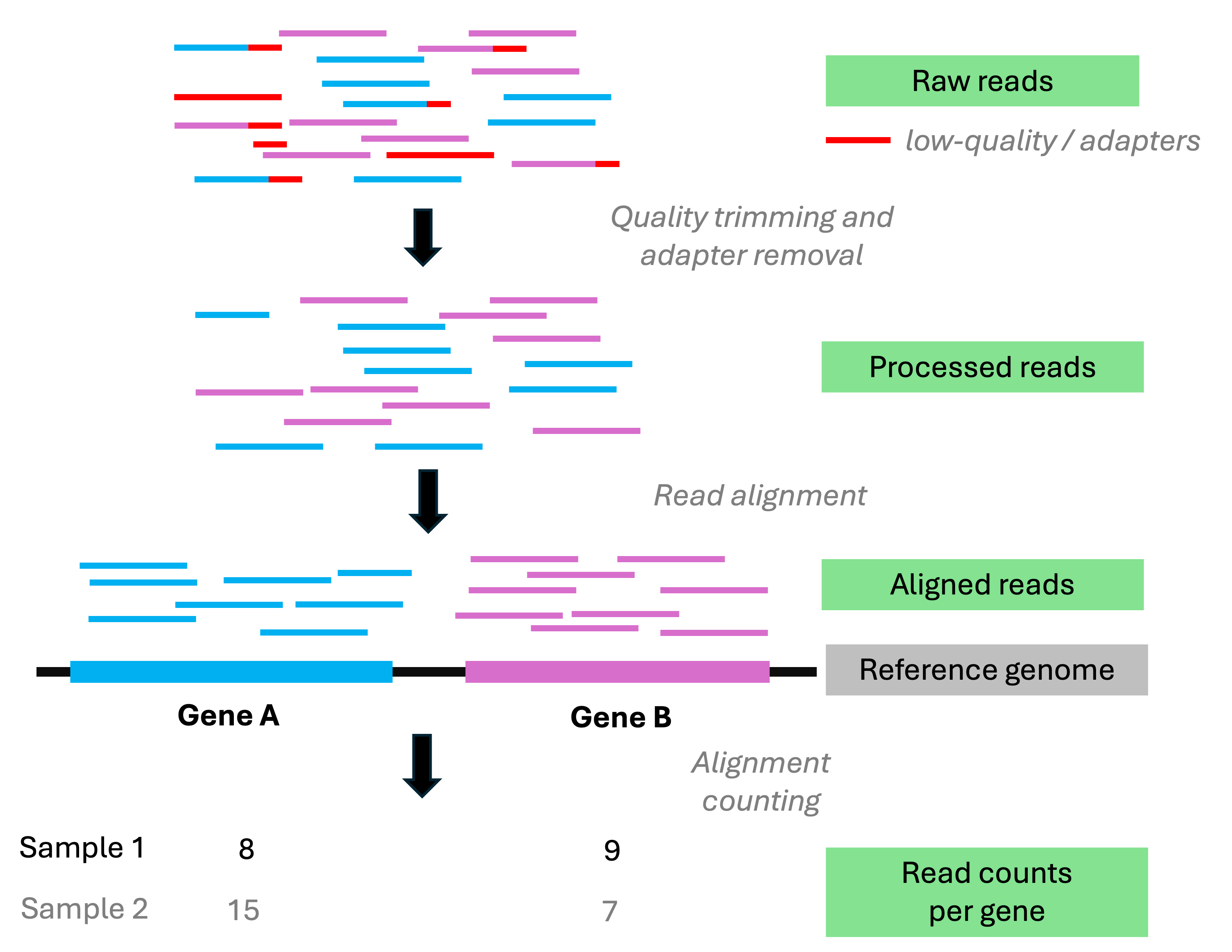
A quick primer on the count table
The count table has one row for each gene and one column for each sample,
with the entries being the number of reads mapping to each gene in each sample:
| Sample 1 | Sample 2 | Sample 3 | |
|---|---|---|---|
| Gene A | 1500 | 2300 | 1800 |
| Gene B | 0 | 5 | 2 |
| Gene C | 300 | 250 | 400 |
Actual count tables have thousands of genes (rows) and usually dozens of samples (columns)
How do you get to such a count table?
A genAI diagram that explains RNA-Seq analysis
I though I’d get some generative AI help with the diagram on the previous slide –
this is what Adobe Firefly came up with: 😵💫
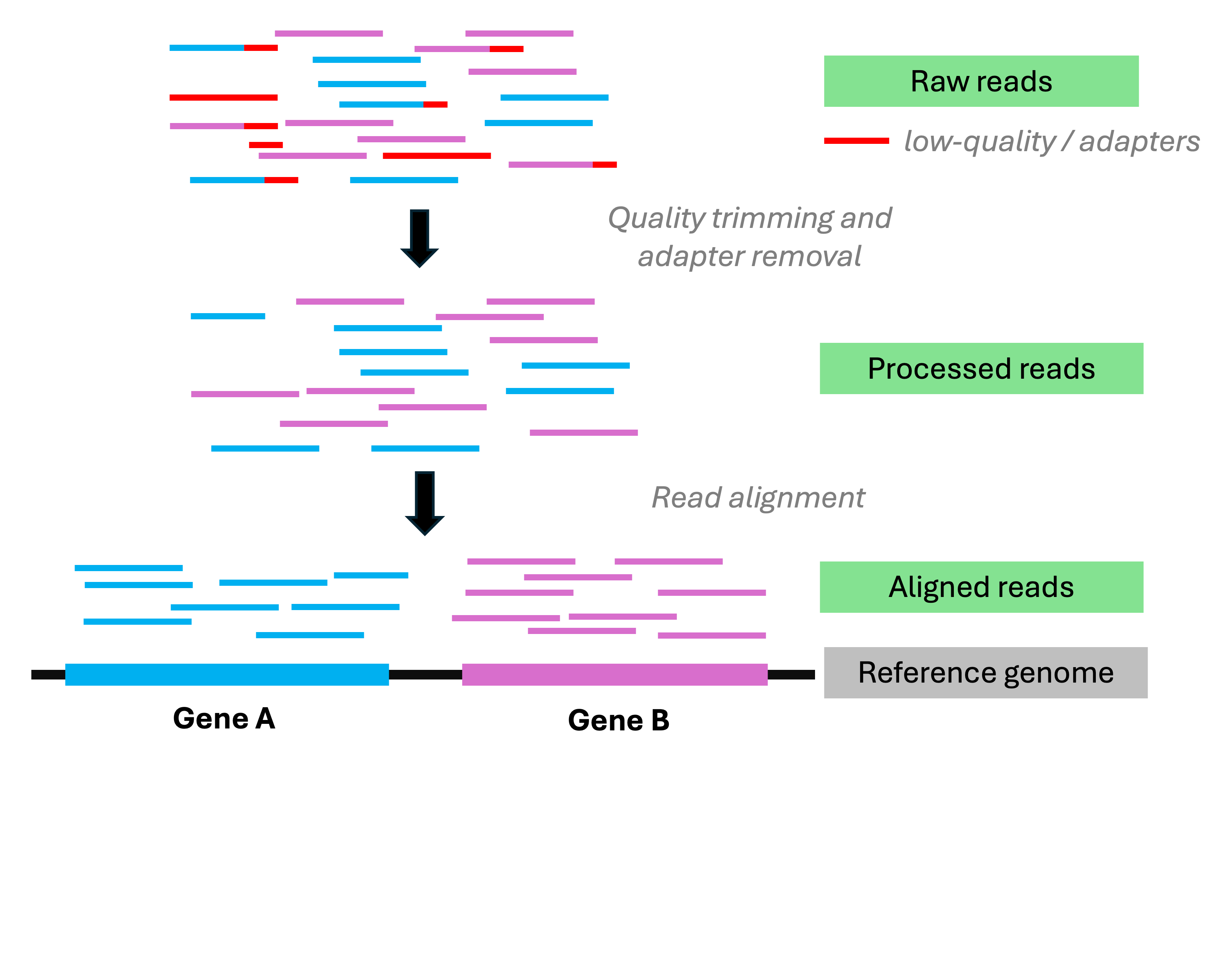
From reads to counts: overview
It specifically involves, at a minimum:
Read preprocessing
Aligning reads to a reference genome
Quantifying expression levels
This part is “bioinformatics”-heavy, with large files, high computing needs, and using command-line tools ran in the Unix shell.
If this is a problem: the process is fairly standardized and suitable to be outsourced.
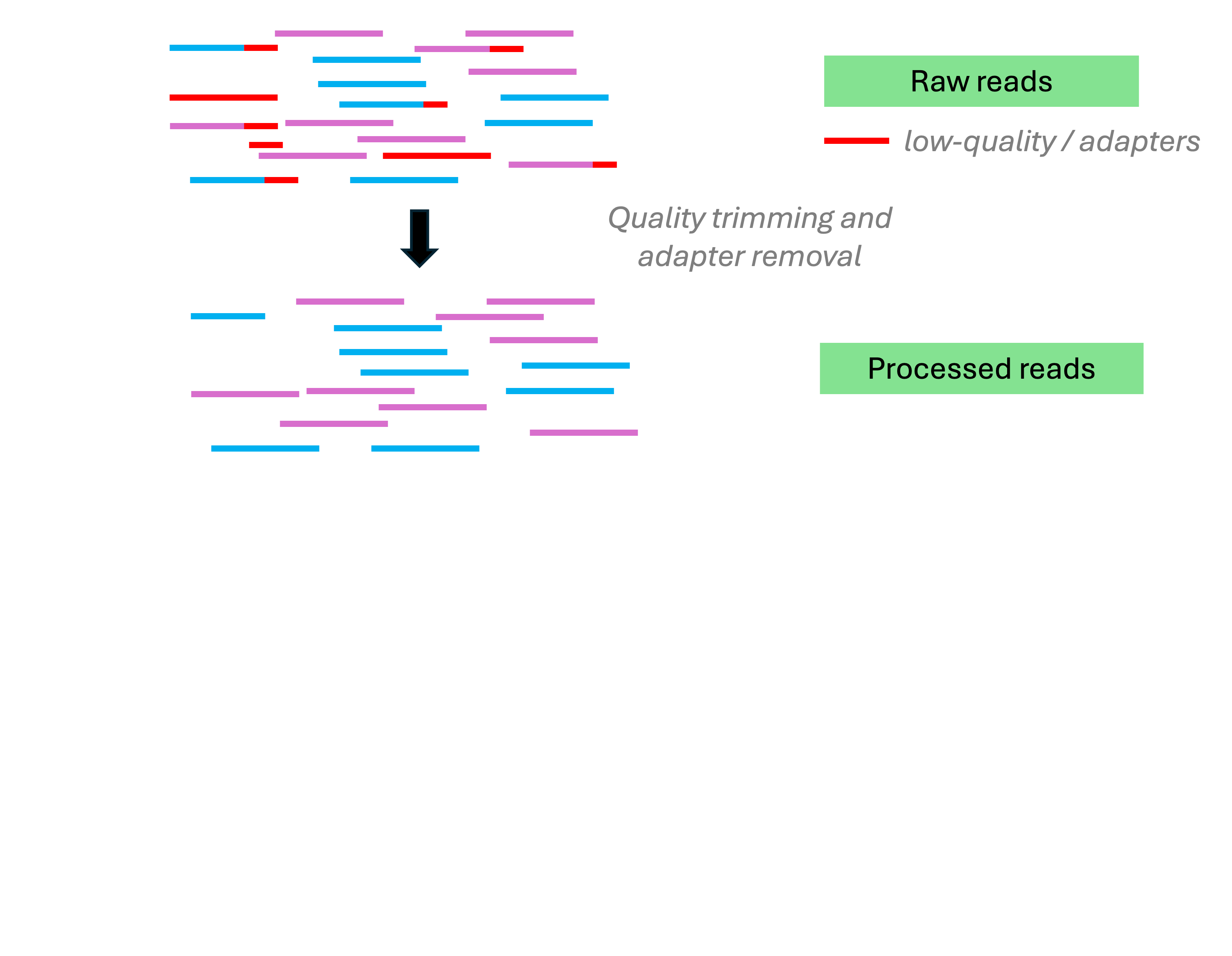
Read pre-processing
Read pre-processing includes:
- Checking the quantity and quality of your reads (e.g. with
FastQC)
- Removing unwanted sequences, such as:
- Adapters, low-quality bases, and very short reads
- rRNA-derived reads (optional)
- Contaminant sequences (optional)
Read alignment to a reference genome
Consider what you know about how mRNAs are produced. Does an RNA transcript always correspond to a contiguous stretch of the genome? If not, what implications does this have for read alignment?
No, it doesn’t, because of splicing. This means that a read can align partially to two different exons, skipping the intron in between. “Regular” genome alignment would not be able to handle this properly – see the next slides.Side note: Splicing
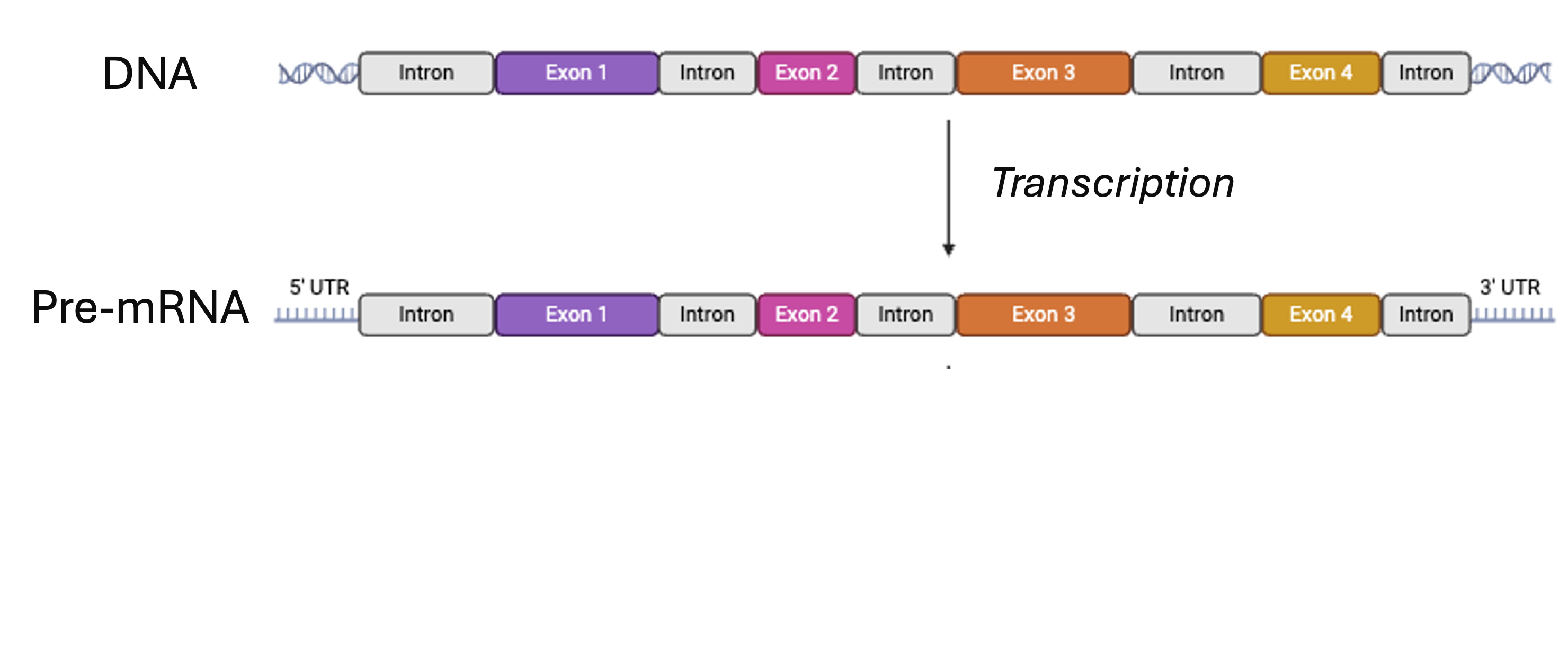
From BioRender
Side note: Splicing
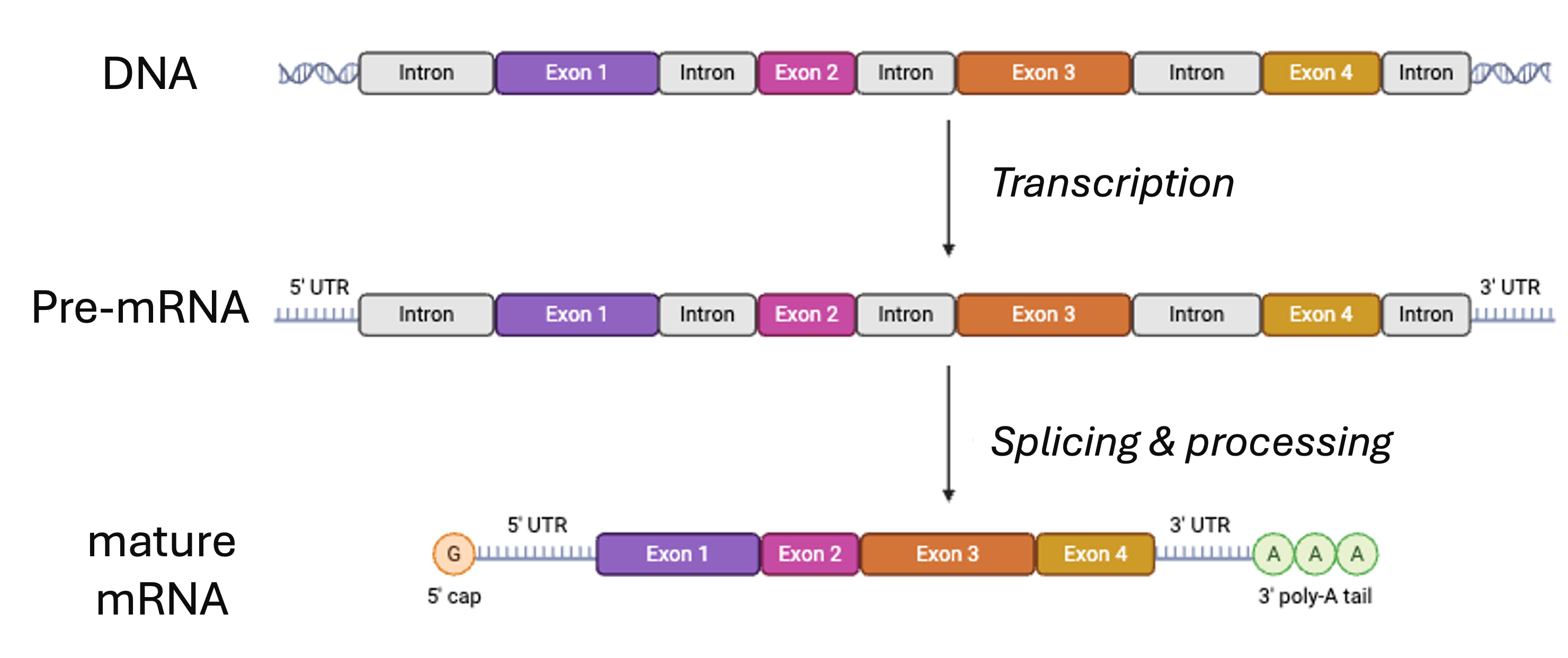
From BioRender
Read alignment to a reference genome
So, the alignment of reads to a reference genome needs to be “splice-aware”:
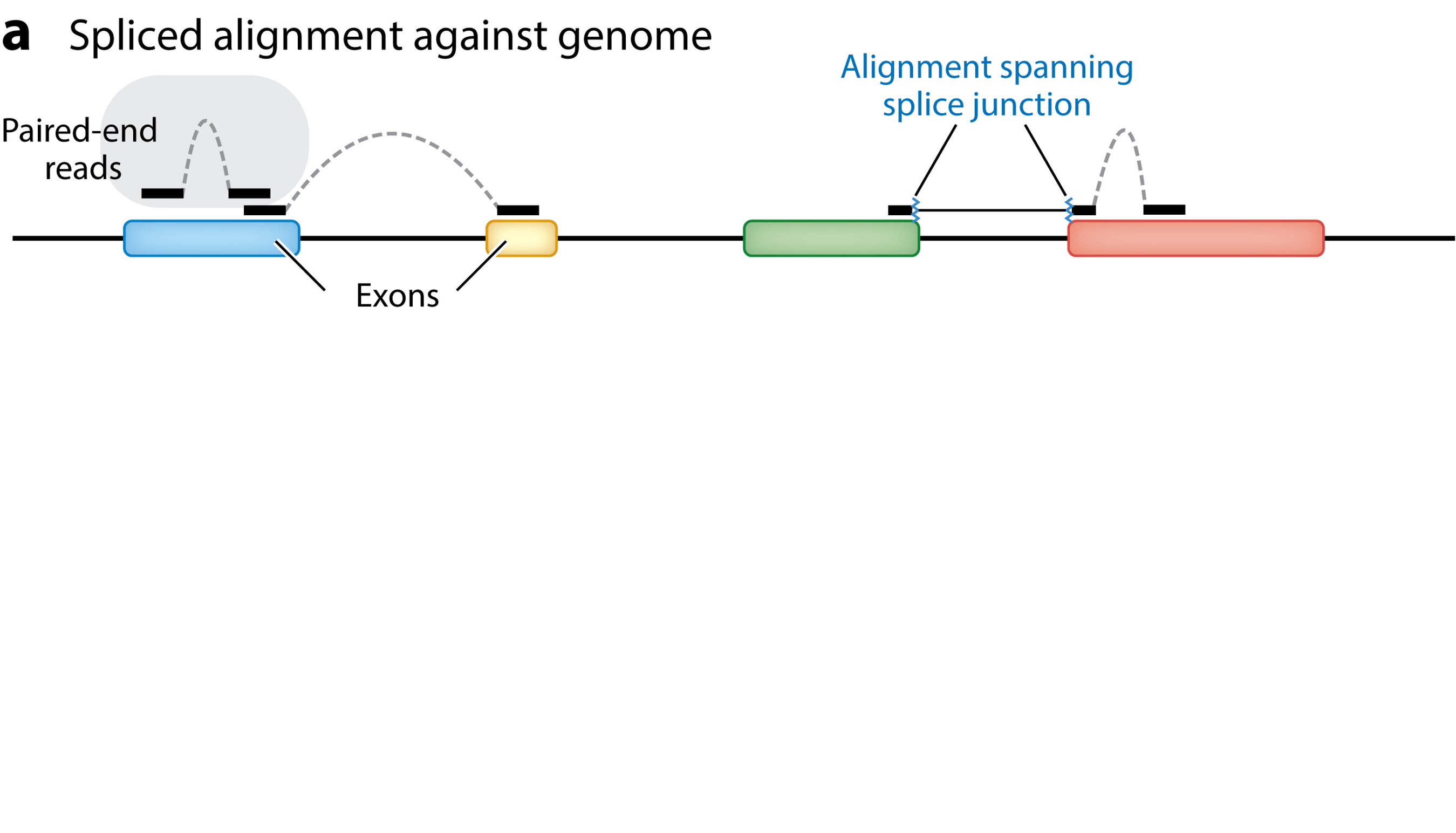
Van den Berge et al. (2019)
Read alignment to a reference genome
Alternatively, you can align to the transcriptome (i.e., all mature transcripts):
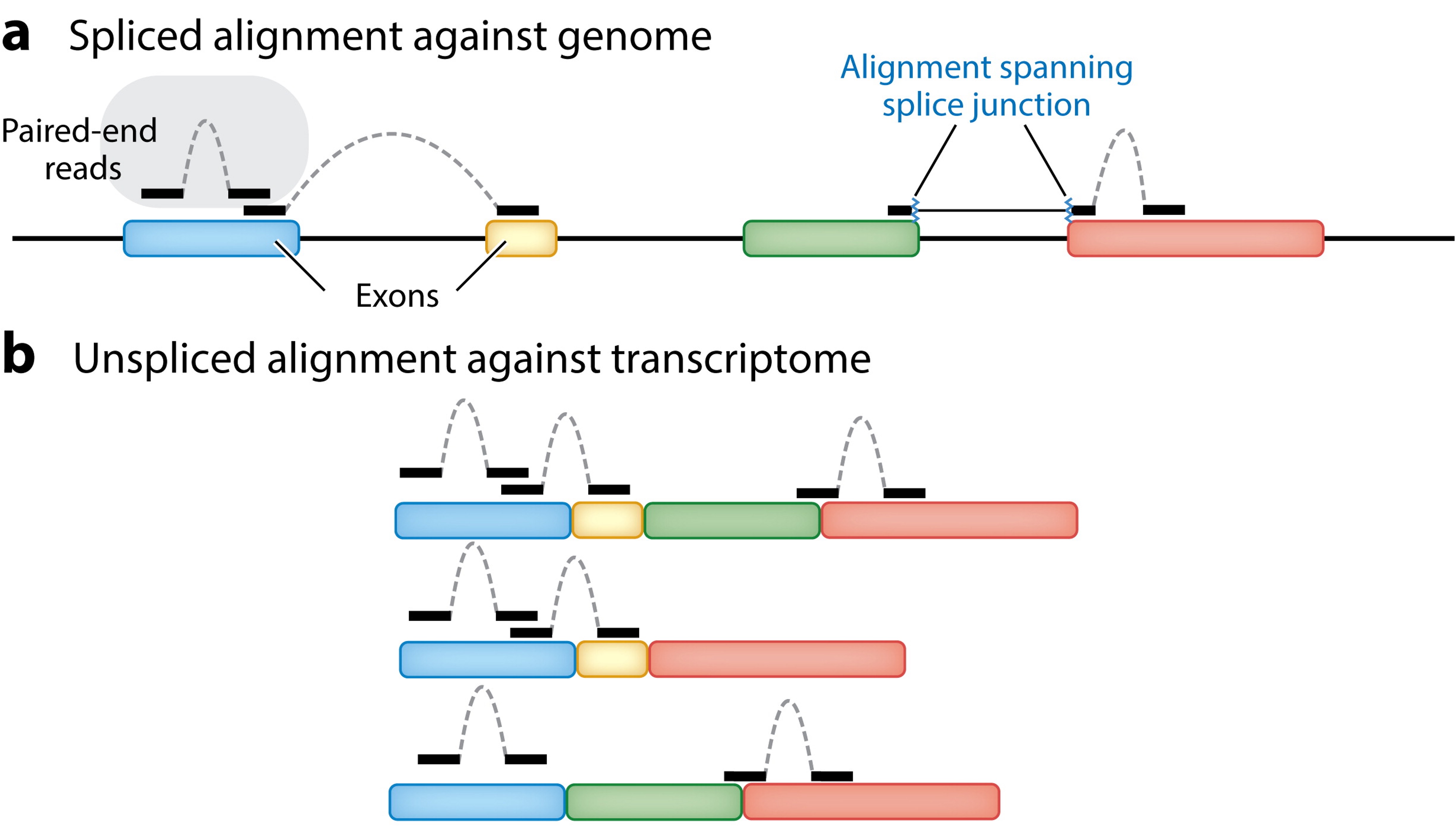
Van den Berge et al. (2019)
Read alignment to a reference genome
Why are there multiple bars in panel b? What do these represent?
These represent different transcripts originating from the same gene due to alternative splicing. These will produce different proteins, which are called isoforms.
Most short-read RNA-Seq studies do not attempt to distinguish between isoforms,
but rather quantify expression at the gene level.

Van den Berge et al. (2019)
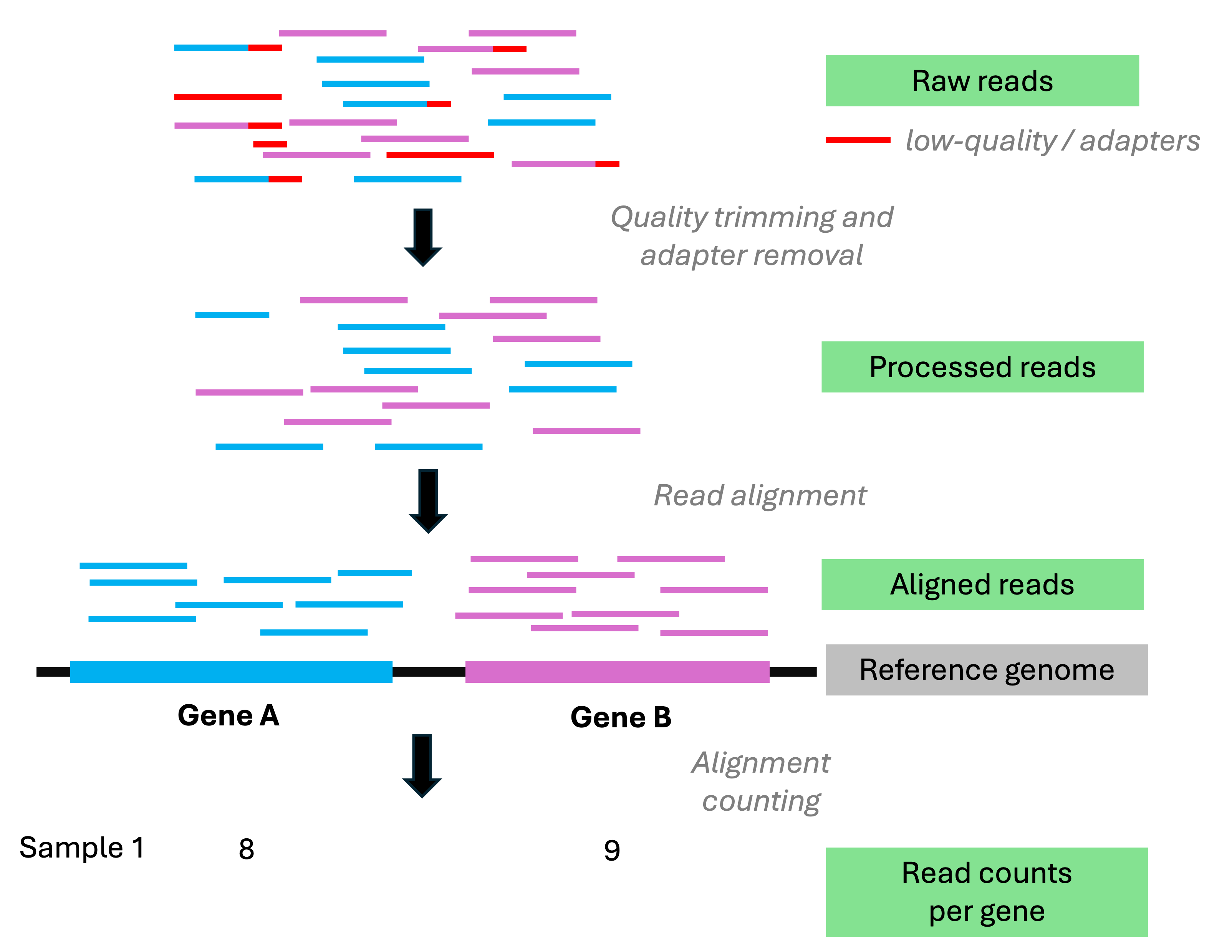
Gene-wise quantification
In essence, a simple counting exercise once you have the alignments in hand:
for each sample, how many reads map to each gene?
Though in practice, a bit more complicated than this, due to e.g.:
- Reads that map to multiple genes (“multi-mapping reads”)
- Sequencing biases and RNA fragmentation
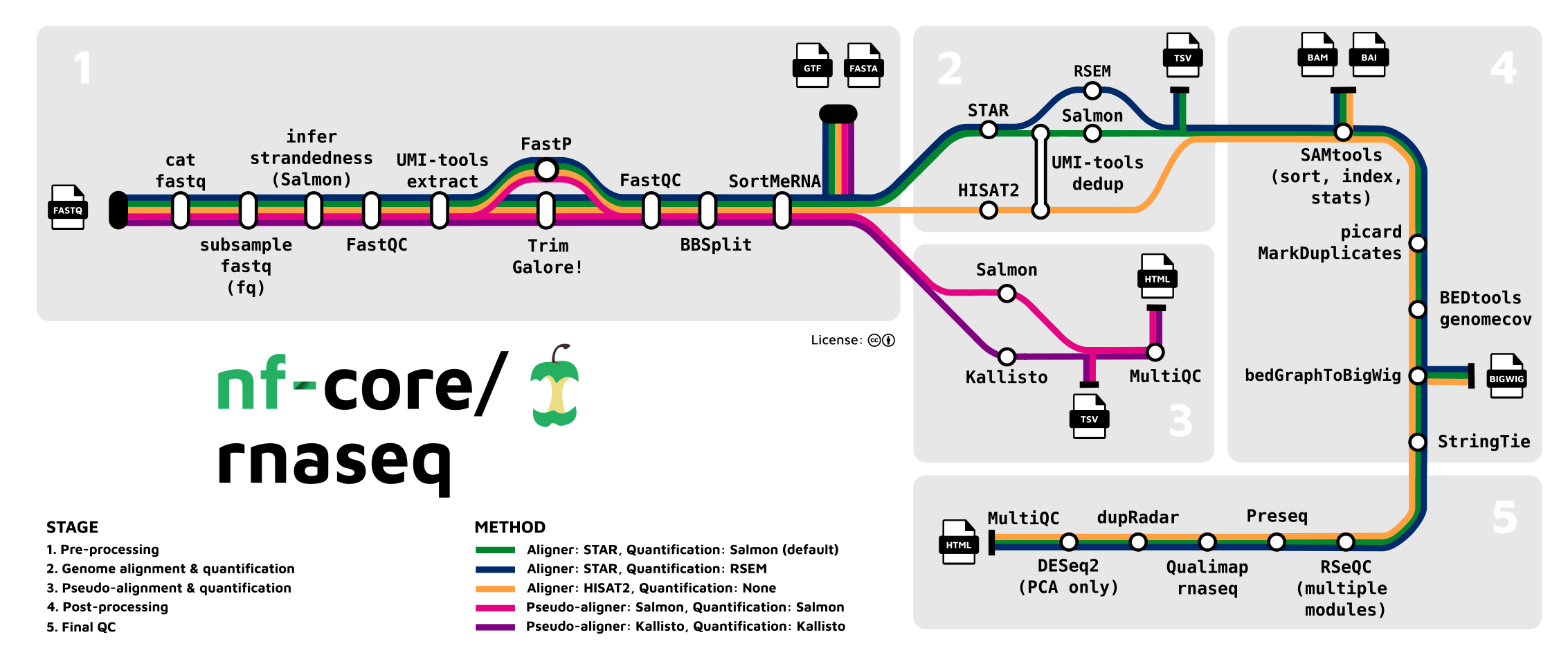
A best-practice pipeline to produce counts
The “nf-core” initiative (https://nf-co.re, Ewels et al. (2020)) aims to produce best-practice and automated bioinformatics pipelines, like for RNA-Seq (https://nf-co.re/rnaseq):
Stage IV: From counts to conclusions
Count table analysis: overview
The second part of RNA-Seq data analysis involves analyzing the count table.
In contrast to the first part, this can be done on a laptop and instead is heavier on
statistics, data visualization and biological interpretation.
It is typically done with the R language, and common aspects include:
Principal Component Analysis (PCA)
Assessing overall sample “clustering” (similarity) patternsDifferential Expression (DE) analysis
Finding genes that differ in expression level between sample groups (DEGs)Functional enrichment analysis
See whether certain gene function categories are overrepresented among DEGs
Principal Component Analysis (PCA)
PCA examines overall patterns of dissimilarity among samples,
such as whether groups of interest form distinct clusters:
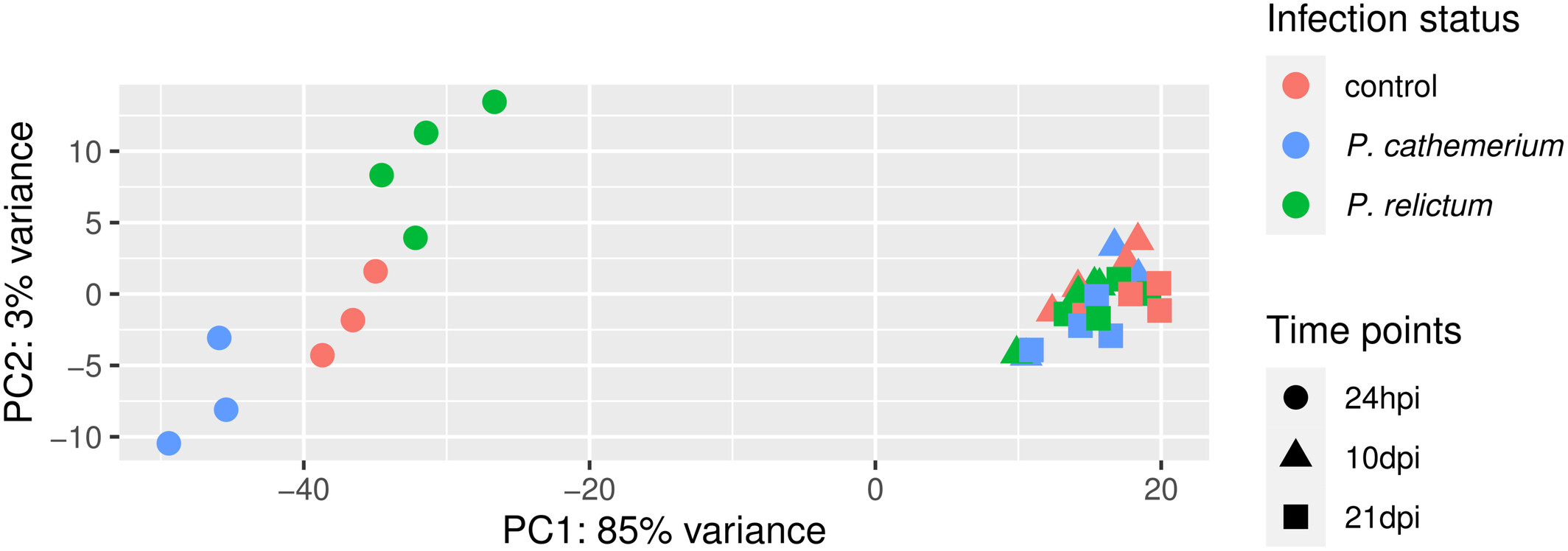
Fig. 1 from Garrigós et al. (2025)
We’ll talk more about the interpretation of this PCA plot in tomorrow’s lab
Principal Component Analysis (PCA)
PCA examines overall patterns of dissimilarity among samples,
such as whether groups of interest form distinct clusters:

Fig. 1 from Garrigós et al. (2025)
PCA is a very useful technique, not just for RNA-Seq data.
To learn more about how it works, see this video (short overview) and this video (more detailed).
Differential expression (DE) analysis
Differential Expression (DE) analysis allows you to test, separately for every expressed gene in your dataset, whether it significantly differs in expression level between groups.
Typically, this is done with pairwise comparisons between groups:
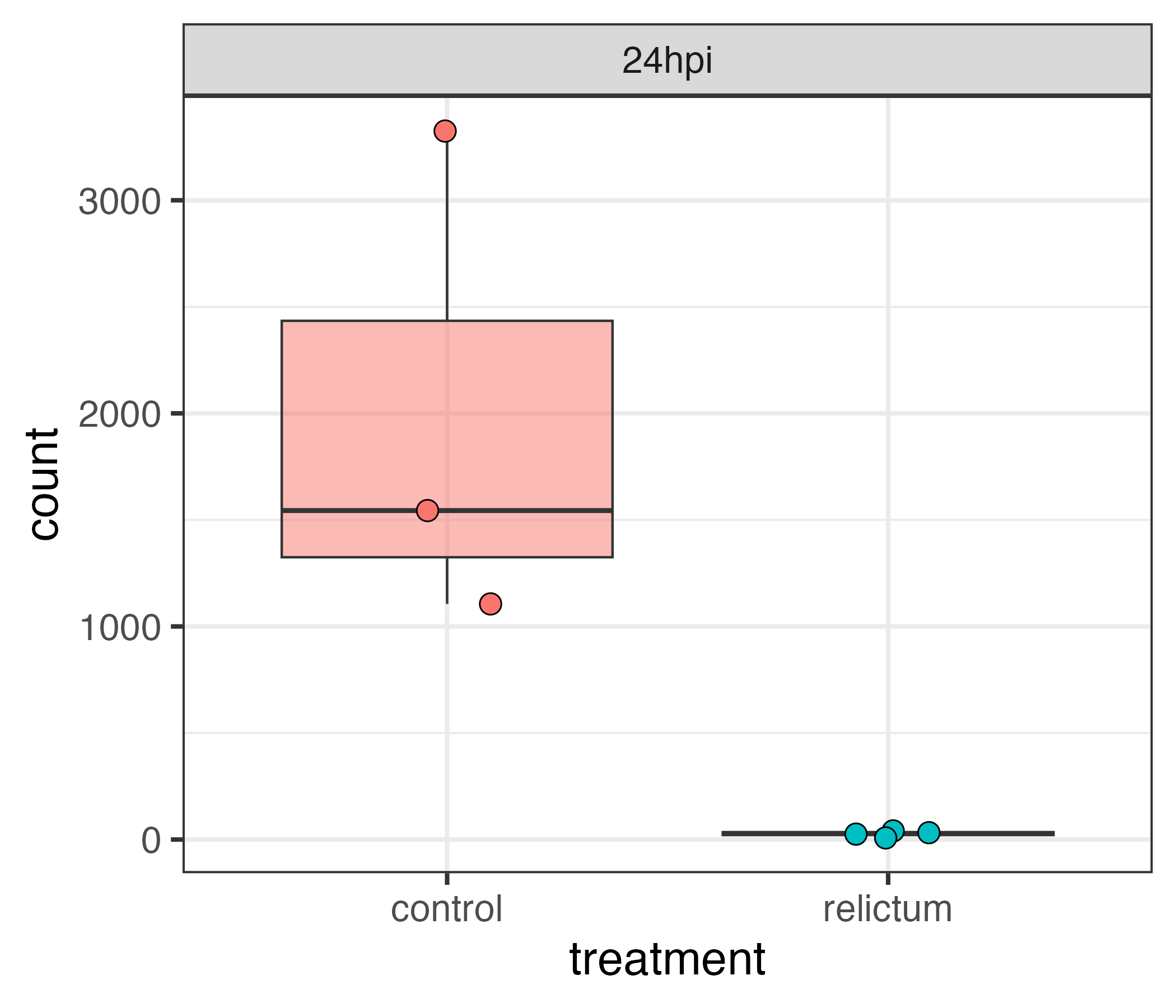
Differential expression (DE) analysis
Differential Expression (DE) analysis allows you to test, separately for every expressed gene in your dataset, whether it significantly differs in expression level between groups.
Typically, this is done with pairwise comparisons between groups:

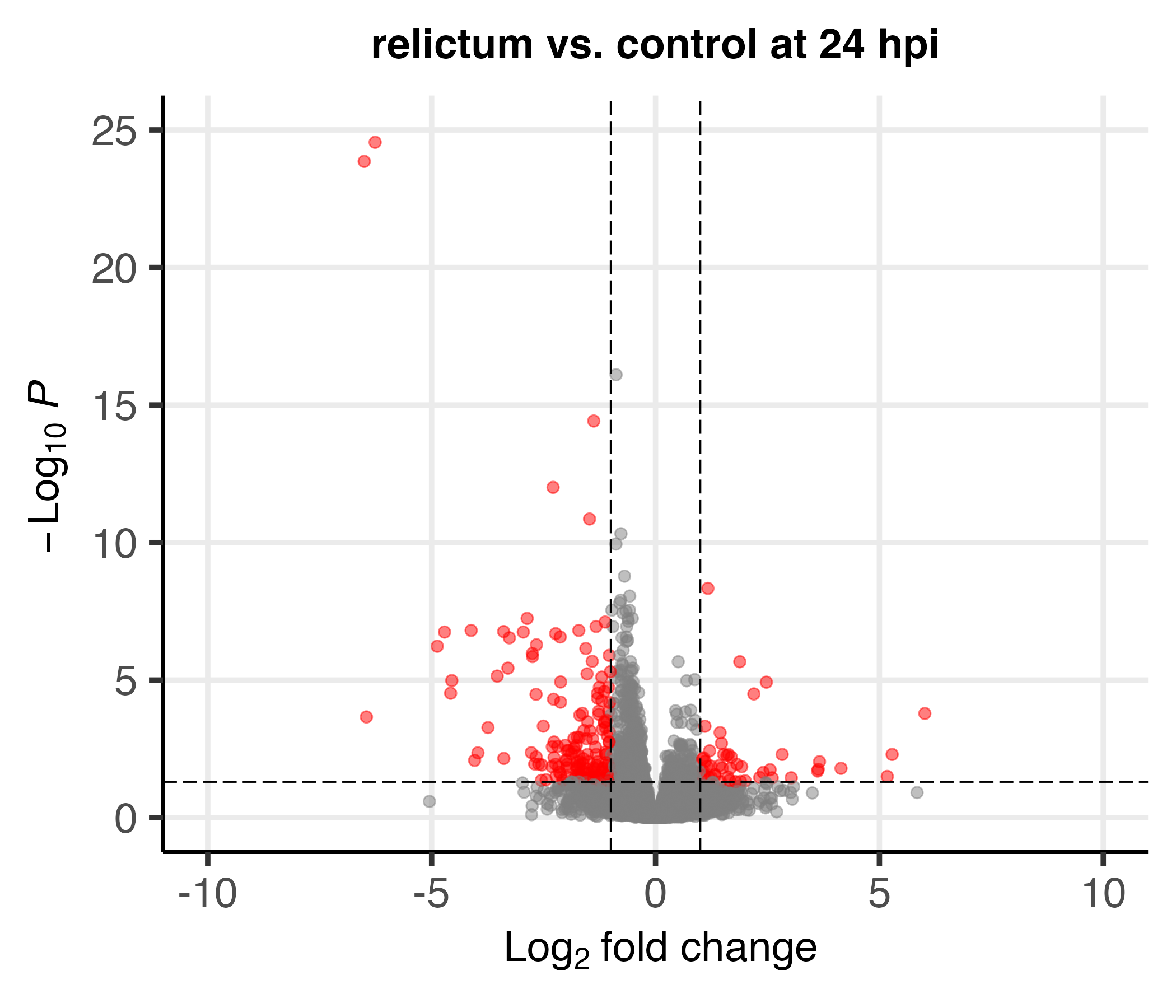
Statistical considerations for the DE analysis
- Gene count normalization
- Probability distribution of the gene count data
- Multiple-testing correction
We will talk about these during tomorrow’s lab.
While not necessarily easy conceptually, this is all fairly straightforward in practice: specialized R packages like DESeq2 take care of the details.
Functional enrichment: introduction
Lists of DEGs can be long and not always easy to make biological sense of.
Functional enrichment analysis helps with this, asking whether certain functional categories of genes are statistically overrepresented among DEGs.
Several databases group genes into such functional categories —
the two main ones used for enrichment analysis are:
- Gene Ontology (GO)
Have a hierarchical structure with more specific terms grouping into more general terms - Kyoto Encyclopedia of Genes and Genomes (KEGG)
Less general than GO, has pathways whose genes can be drawn and connected in diagrams
Functional enrichment: conceptually
A GO term called “photosynthesis” is associated with a set of genes involved in photosynthesis.
Say that 300 genes out of 30,000 genes in the genome have been annotated to this term = 1%
Say that among your 50 out of your 500 DEGs are annotated to this term = 10%
The larger the difference between these percentages, the stronger the indication that the function in question –here, photosynthesis– is overrepresented among your DEGs.
Functional enrichment: GO
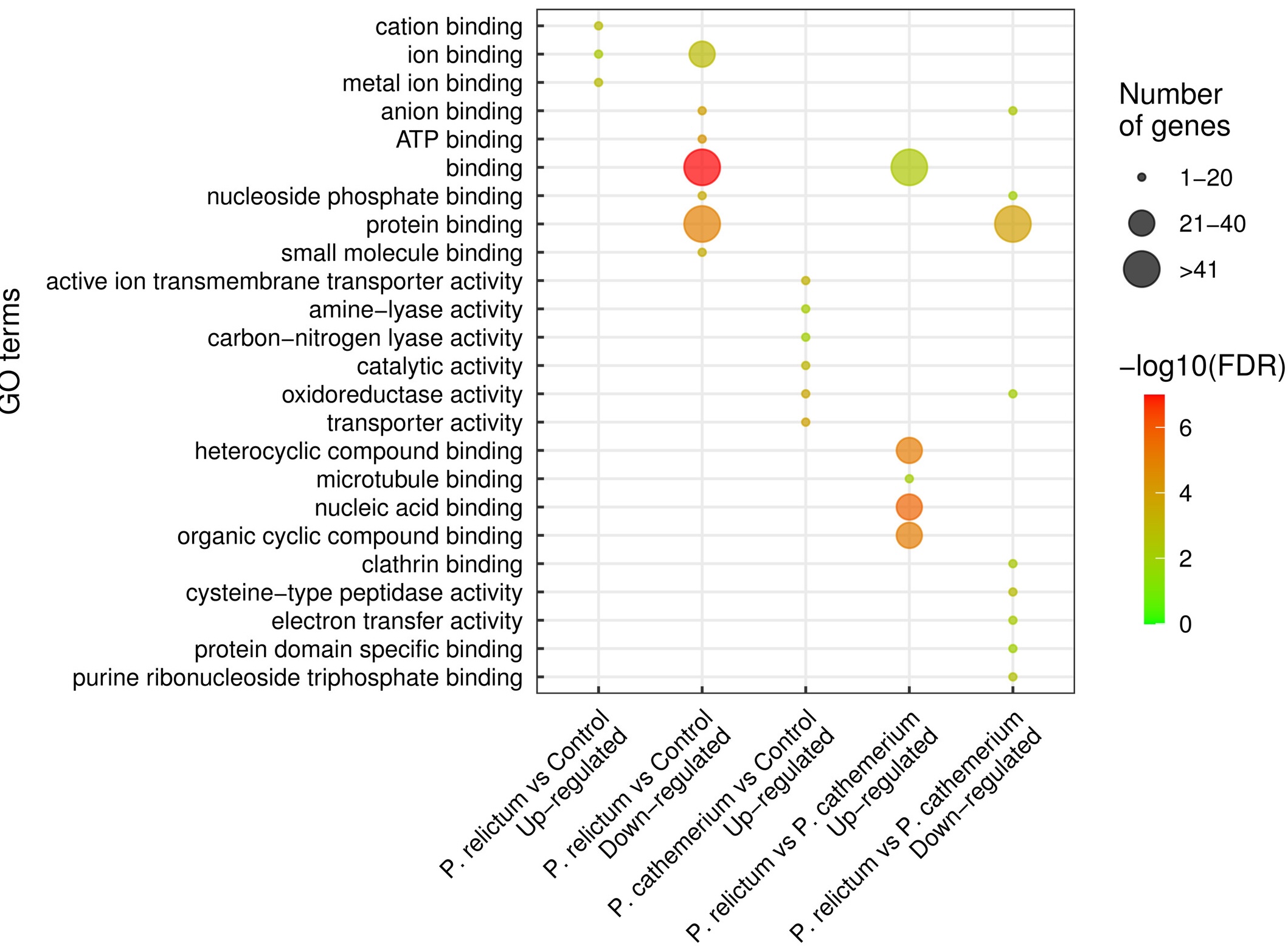
Fig. 4 from Garrigós et al. (2025)
Functional enrichment: KEGG
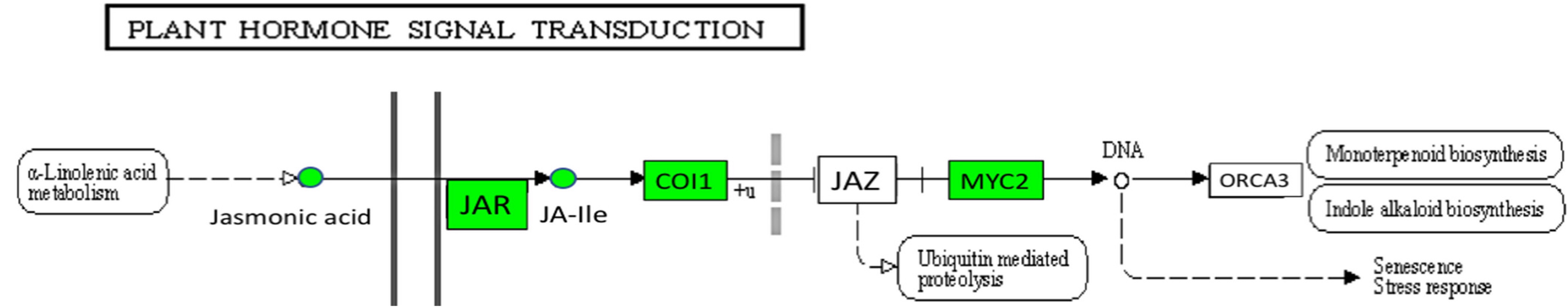
Rodriguez et al. (2020)
KEGG representation of up-regulated genes related to jasmonic acid (JA) signal transduction pathways (ko04075) in banana cv. Calcutta 4 after inoculation with Pseudocercospora fijiensis. Genes or chemicals up-regulated at any time point were highlighted in green.
Questions?
Bonus: Read alignment QC
- Alignment rates
What percentage of reads was successfully aligned? (Should be >80%)
- Alignment targets
What percentages of aligned reads mapped to exons vs. introns vs. intergenic regions?